Gabriel Machado

My personal portfolio
Welcome to Gabriel-Machado-GM.GITHUB.IO -> My Personal Portfolio
Gabriel Machado - Data Scientist
Skills: Python, SQL, R, Excel, AWS, Big Data Tools (Google Data Studio/Looker, Zoho Analytics, Tableau, Power BI)
Contact: (011) 95459-2057 | gabriel.machado.gmbr@gmail.com | LinkedIn | GitHub/Portfolio
Objective
Seeking opportunities in Data Science with a focus on Extract, Transform, Load (ETL), Data Analysis, Retrieval Augmented Generation (RAG), Business Intelligence, Machine Learning, and architecting AWS solutions.
Education
-
B.S., Data Science and Artificial Intelligence UNIVESP (Universidade Virtual do Estado de São Paulo) (Expected Graduation: 2025)
Relevant Knowledge & Skills
- Python (Advanced): Object-Oriented Programming (OOP), modularization, file handling, data cleaning, exploratory analysis, supervised (regression, classification) and unsupervised (recommendations, reinforcement learning) machine learning, advanced algorithms.
- Libraries: Scikit-Learn, NumPy, Pandas, Matplotlib, Plotly, Docling, PyTorch, LangChain, MarkItDown.
- Certification: Stanford University & DeepLearning.AI.
- SQL (Advanced): Complex query optimization, function creation, Entity-Relationship Diagrams (ERD), database modeling (relational, key-value, document-oriented, graph).
- R (Advanced): Exploratory data analysis (conditional probability, inference, estimation, hypothesis testing, sampling distributions), data cleaning, processing, and visualization.
- Certification: “The Data Scientist’s Toolbox” - Johns Hopkins University.
- Excel (Advanced): Data organization, filtering, sorting, functions/formulas for cleaning and transformation, statistical/mathematical analysis, charts, pivot tables, macro creation/editing, VBA, scenario analysis, forecasting, interactive dashboards, collaboration.
- Certification: “Excel Skills for Business” - Macquarie University.
- Amazon Web Services (AWS): Practical labs and concepts including EC2, S3, Lambda, RDS, VPC, IAM (Roles, Policies), CloudFront, ELB, CloudWatch, SNS, Route 53. Developed soft skills like Active Listening, Time Management, Effective Communication, and Conflict Management through AWS Re/Start program.
- Certification: AWS Certified Cloud Practitioner.
- Other Big Data Tools: Experience with Google Data Studio / Looker, Zoho Analytics, Tableau, and Power BI.
Work Experience
AI Engineer @ G.A.S. Global Actuarial Solution (2023 - Present)
- Applied AI in Actuarial Expertise. Managed calculation engine parameters (developed in Excel), monitored bugs, implemented improvements, and created auxiliary functions using SQL/Python.
- Innovated by creating a database of legal proceedings (PDFs converted to Markdown) and built Retrieval Augmented Generation (RAG) systems.
- Implemented a complete pipeline for the “Actuarial Expertise” process using Python and libraries like Docling, PyTorch, LangChain, MarkItDown.
Business Intelligence Specialist @ Farol Do Futuro (2020 - 2023)
- Acted as a BI Specialist for 3 years, implementing automated dashboards and reports.
- Analyzed data to optimize processes and strategic decisions.
- Managed databases and KPIs.
- Developed predictive models to identify growth opportunities.
- Collaborated with teams to transform data into actionable insights.
Apprentice Electrical Technician @ Enel Brasil S.A. (2019)
- Performed dimensioning calculations for residential and industrial electrical networks using Excel.
- Created diagrams and projects.
- Studied and applied NBR (Brazilian standards) and NR (regulatory standards).
Languages
- English: C1 Advanced (CEFR) - EF SET Certified (Note: Replace YOUR_CERTIFICATE_ID with the actual ID from the certificate if available, otherwise remove the link)
- Spanish: Intermediate
Volunteer Work
ACM - Alphaville (2016 - 2017)
- Monitor/Collaborator in 12 major events.
- Vice-President of the Leaders’ Body (2017).
- Attended São Paulo Leaders Meeting (2017).
- Attended National Leaders Meeting (2017).
Projects
Data-Driven EEG Band Discovery with Decision Trees
Developed objective strategy for discovering optimal EEG bands based on signal power spectra using Python. This data-driven approach led to better characterization of the underlying power spectrum by identifying bands that outperformed the more commonly used band boundaries by a factor of two. The proposed method provides a fully automated and flexible approach to capturing key signal components and possibly discovering new indices of brain activity.
Decoding Physical and Cognitive Impacts of Particulate Matter Concentrations at Ultra-Fine Scales
Used Matlab to train over 100 machine learning models which estimated particulate matter concentrations based on a suite of over 300 biometric variables. We found biometric variables can be used to accurately estimate particulate matter concentrations at ultra-fine spatial scales with high fidelity (r2 = 0.91) and that smaller particles are better estimated than larger ones. Inferring environmental conditions solely from biometric measurements allows us to disentangle key interactions between the environment and the body.
Talks & Lectures
- Causality: The new science of an old question - GSP Seminar, Fall 2021
- Guest Lecture: Dimensionality Reduction - Big Data and Machine Learning for Scientific Discovery (PHYS 5336), Spring 2021
- Guest Lecture: Fourier and Wavelet Transforms - Scientific Computing (PHYS 5315), Fall 2020
- A Brief Introduction to Optimization - GSP Seminar, Fall 2019
- Weeks of Welcome Poster Competition - UTD, Fall 2019
-
A Brief Introduction to Networks - GSP Seminar, Spring 2019
- Data Science YouTube
Publications
- Talebi S., Lary D.J., Wijeratne L. OH., and Lary, T. Modeling Autonomic Pupillary Responses from External Stimuli Using Machine Learning (2019). DOI: 10.26717/BJSTR.2019.20.003446
- Wijeratne, L.O.; Kiv, D.R.; Aker, A.R.; Talebi, S.; Lary, D.J. Using Machine Learning for the Calibration of Airborne Particulate Sensors. Sensors 2020, 20, 99.
- Lary, D.J.; Schaefer, D.; Waczak, J.; Aker, A.; Barbosa, A.; Wijeratne, L.O.H.; Talebi, S.; Fernando, B.; Sadler, J.; Lary, T.; Lary, M.D. Autonomous Learning of New Environments with a Robotic Team Employing Hyper-Spectral Remote Sensing, Comprehensive In-Situ Sensing and Machine Learning. Sensors 2021, 21, 2240. https://doi.org/10.3390/s21062240
- Zhang, Y.; Wijeratne, L.O.H.; Talebi, S.; Lary, D.J. Machine Learning for Light Sensor Calibration. Sensors 2021, 21, 6259. https://doi.org/10.3390/s21186259
- Talebi, S.; Waczak, J.; Fernando, B.; Sridhar, A.; Lary, D.J. Data-Driven EEG Band Discovery with Decision Trees. Preprints 2022, 2022030145 (doi: 10.20944/preprints202203.0145.v1).
- Fernando, B.A.; Sridhar, A.; Talebi, S.; Waczak, J.; Lary, D.J. Unsupervised Blink Detection Using Eye Aspect Ratio Values. Preprints 2022, 2022030200 (doi: 10.20944/preprints202203.0200.v1).
- Talebi, S. et al. Decoding Physical and Cognitive Impacts of PM Concentrations at Ultra-fine Scales, 29 March 2022, PREPRINT (Version 1) available at Research Square [https://doi.org/10.21203/rs.3.rs-1499191/v1]
- Lary, D.J. et al. (2022). Machine Learning, Big Data, and Spatial Tools: A Combination to Reveal Complex Facts That Impact Environmental Health. In: Faruque, F.S. (eds) Geospatial Technology for Human Well-Being and Health. Springer, Cham. https://doi.org/10.1007/978-3-030-71377-5_12
- Wijerante, L.O.H. et al. (2022). Advancement in Airborne Particulate Estimation Using Machine Learning. In: Faruque, F.S. (eds) Geospatial Technology for Human Well-Being and Health. Springer, Cham. https://doi.org/10.1007/978-3-030-71377-5_13
"""
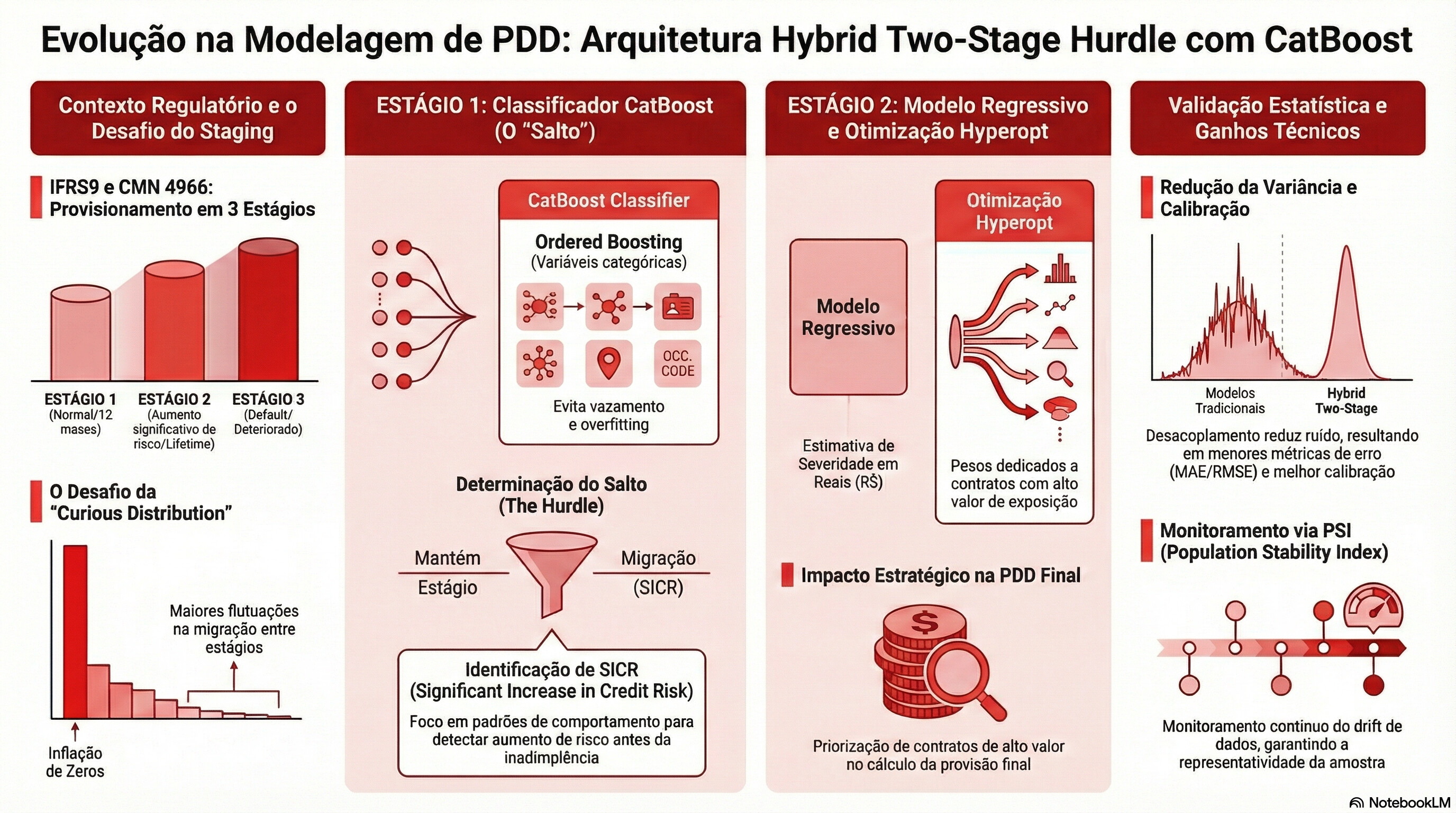
IFRS 9 Multi-Strategy Sampler Suite — Static Samples for Experimentation
=========================================================================
Generates multiple static Delta-persisted samples from the full IFRS 9
provision output table, each designed to isolate a different analytical
dimension for downstream ``model_experimentation_suite`` runs.
Samples Generated:
A1 Neyman 30-Strata (Stage × LGD Decil) → ``tb_spl_ney_decil_<cart>``
A2 Neyman by Stage × id_lgd_segment → ``tb_spl_ney_lgdseg_<cart>``
A3 Neyman by Stage × ead_cli → ``tb_spl_ney_ead_<cart>``
B Migration-Active Oversampling → ``tb_spl_migr_actv_<cart>``
C-E Stage-Isolated (Stage 1, 2, 3) → ``tb_spl_stg{n}_<cart>``
Validation:
Every sample is validated against the source population using:
- PSI (Population Stability Index) — classic Lewis thresholds
- PRS (Population Resemblance Statistic) — dynamic Chi-Square bounds
Enterprise Databricks PySpark Standards:
- Data Primitives: EXCLUSIVELY ``pyspark.sql.functions``.
- Pandas UDFs: PROHIBITED. No ``.toPandas()`` for computation.
- Storage: All outputs in Delta format.
- Documentation: pdDoc standard on every function.
Author: Principal Data Engineer & ML Architect
"""
from __future__ import annotations
import gc
import math
import re
from datetime import datetime
from dataclasses import dataclass, field
from typing import Any, Dict, List, Optional, Tuple
from pyspark.sql import SparkSession, DataFrame, Window
import pyspark.sql.functions as F
from pyspark import StorageLevel
from pyspark.sql.types import (
StructType, StructField, StringType, DoubleType, IntegerType, LongType,
)
from neyman_rnabox_sampler import RNABOXAllocator
from prs_validator import PRSValidator
# ╔═════════════════════════════════════════════════════════════════════════╗
# ║ 1. CONFIGURATION ║
# ╚═════════════════════════════════════════════════════════════════════════╝
# ── Column Name Lists (Organised by Semantic Type) ──
# Edit ONLY these three lists. DICT_CASTINGS is auto-generated below.
# Each list drives automatic type casting:
# COLS_FLAGS → cast to "int" (binary 0/1 indicators)
# COLS_NUMERICS → cast to "double" (continuous measures)
# COLS_CATEGORICALS → cast to "string" (categorical dimensions)
#
# Additional fixed columns (stage → int, reporting_date → date) are
# appended automatically and should NOT be placed in these lists.
COLS_FLAGS: List[str] = [
# ── Binary Indicators (0/1) ──
# Add flag column names here, e.g.:
# "flag_reestruturado",
# "flag_default_anterior",
]
COLS_NUMERICS: List[str] = [
# ── Continuous / Numeric Measures ──
"output_provision_amount",
"lgd_pond",
"ead_cli",
"pd_12m",
"exposure",
"lgd_estimate",
"ead",
"days_past_due",
"remaining_maturity_months",
"interest_rate",
"collateral_coverage_ratio",
]
COLS_CATEGORICALS: List[str] = [
# ── Categorical / String Dimensions ──
"carteira_ifrs",
"carteira_local",
"id_lgd_segment",
"setor_economico",
"rating_bucket",
"contract_id_orig",
]
# ── Auto-Generated Casting Dictionary ──
# Built from the three lists above + fixed structural columns.
# DO NOT edit this dict directly — edit the lists above instead.
def _build_castings(
flags: List[str],
numerics: List[str],
categoricals: List[str],
) -> Dict[str, str]:
"""Builds the unified casting dictionary from semantic column lists.
Args:
flags (List[str]): Binary indicator column names.
numerics (List[str]): Continuous/numeric column names.
categoricals (List[str]): Categorical/string column names.
Expected Input Schema:
Three plain lists of column name strings.
Output Schema Transformation:
Returns Dict[str, str] mapping column name → Spark SQL type.
Memory Implications:
Negligible. Dict construction on driver.
Exception Handling Protocols:
Duplicate column names across lists are silently deduplicated
(last-write wins).
Returns:
Dict[str, str]: Column-name → Spark-type mapping.
"""
castings: Dict[str, str] = {}
for col_name in flags:
castings[col_name] = "int"
for col_name in numerics:
castings[col_name] = "double"
for col_name in categoricals:
castings[col_name] = "string"
# Fixed structural columns (always present)
castings["stage"] = "int"
castings["reporting_date"] = "date"
return castings
DICT_CASTINGS: Dict[str, str] = _build_castings(
COLS_FLAGS, COLS_NUMERICS, COLS_CATEGORICALS,
)
@dataclass
class SamplerConfig:
"""
Centralised, immutable configuration for the IFRS 9 Sampler Suite.
Attributes:
spark (SparkSession): Active Spark session.
source_table (str): Fully-qualified source table name.
dest_schema (str): Unity Catalog schema for output tables.
carteira_ifrs (str): Portfolio filter value.
reporting_date_start (Optional[str]): ISO date string
(e.g., ``"2022-01-01"``) to filter ``reporting_date >=``.
If None, no temporal lower-bound filter is applied.
target_col (str): Column whose variance drives Neyman allocation.
decil_col (str): Column used to build LGD-based deciles.
stage_col (str): IFRS 9 Stage column.
total_sample_size (int): Budget for Neyman samples.
min_stratum_size (int): RNABOX floor constraint per stratum.
confidence_level (float): Z-level for minimum sample size calc.
margin_of_error (float): Desired precision for sample size calc.
dq_chunk_size (int): Number of columns processed per chunk in
the Data Quality Profiler. Controls OOM risk on wide tables.
dq_missing_threshold (float): Maximum missing percentage (0-1)
allowed for a column to survive DQ-based feature selection.
Columns exceeding this in a given stage are dropped.
dq_top_n_categorical (int): Number of top-frequency values to
collect for categorical columns in the Profiler.
psi_n_bins (int): Number of bins for PSI calculation.
prs_epsilon (float): PRS ε-resemblance tolerance.
prs_alpha (float): PRS significance level.
migration_oversample_ratio (float): Oversampling factor for
contracts that migrated stage relative to stable contracts.
trailing_months (int): Lookback window for temporal filtering.
validation_cols (List[str]): Extra columns to validate via PSI/PRS
beyond the target column.
"""
spark: Any = None
source_table: str = "prd.s_stbr_dri_ifr.tb_output_modellica_ifrs9_cred"
dest_schema: str = "prd.sand_crc_estudos_ifrs9"
carteira_ifrs: str = "Corporate"
reporting_date_start: Optional[str] = "2022-01-01"
target_col: str = "output_provision_amount"
decil_col: str = "lgd_pond"
stage_col: str = "stage"
total_sample_size: int = 50_000
min_stratum_size: int = 500
confidence_level: float = 0.95
margin_of_error: float = 0.05
dq_chunk_size: int = 50
dq_missing_threshold: float = 0.40
dq_top_n_categorical: int = 10
psi_n_bins: int = 10
prs_epsilon: float = 0.05
prs_alpha: float = 0.05
migration_oversample_ratio: float = 5.0
trailing_months: int = 12
validation_cols: List[str] = field(
default_factory=lambda: ["lgd_pond", "ead_cli"],
)
# ── Vectorized Column Detection & Explosion ──
vectorized_separator: str = "#"
vectorized_min_pct: float = 0.50
vectorized_numeric_threshold: float = 0.90
vectorized_chunk_size: int = 30
vectorized_explode_chunk_size: int = 5
vectorized_max_elements: Optional[int] = None
vectorized_max_subcols_per_chunk: int = 100
enable_vectorized_explosion: bool = True
# ── Semantic Re-Typing (Fase 3) ──
string_to_numeric_threshold: float = 0.95
flag_max_distinct: int = 5
flag_known_values: Tuple[str, ...] = (
"0", "1", "s", "n", "sim", "nao", "não",
"true", "false", "t", "f", "y", "yes", "no",
)
date_sample_formats: Tuple[str, ...] = (
"yyyy-MM-dd", "dd/MM/yyyy", "yyyy/MM/dd",
"dd-MM-yyyy", "yyyyMMdd",
)
sample_values_count: int = 5
# ── Cluster / Memory Tuning ──
n_repartition: Optional[int] = None
storage_level: str = "MEMORY_AND_DISK"
enable_sample_collection: bool = True
skip_date_inference: bool = False
# ── Delta Checkpointing ──
enable_delta_checkpoints: bool = True
checkpoint_prefix: str = "chkpt_"
# ── Manual Type Hints (Pre-configuração opcional de colunas) ──
manual_numeric_cols: List[str] = field(default_factory=list)
manual_categorical_cols: List[str] = field(default_factory=list)
manual_flag_cols: List[str] = field(default_factory=list)
manual_vectorized_numeric_cols: List[str] = field(default_factory=list)
manual_vectorized_categorical_cols: List[str] = field(default_factory=list)
manual_date_cols: List[str] = field(default_factory=list)
# ╔═════════════════════════════════════════════════════════════════════════╗
# ║ 2. MINIMUM SAMPLE SIZE CALCULATOR (COCHRAN + STRATIFIED) ║
# ╚═════════════════════════════════════════════════════════════════════════╝
def compute_minimum_sample_size(
df: DataFrame,
target_col: str,
stage_col: str,
confidence_level: float = 0.95,
margin_of_error: float = 0.05,
) -> Dict[str, Any]:
"""
Computes the probabilistic minimum sample size required to represent
the population with statistical rigour, using Cochran's formula for
continuous variables with finite-population correction (FPC) and
stratified adjustments per IFRS Stage.
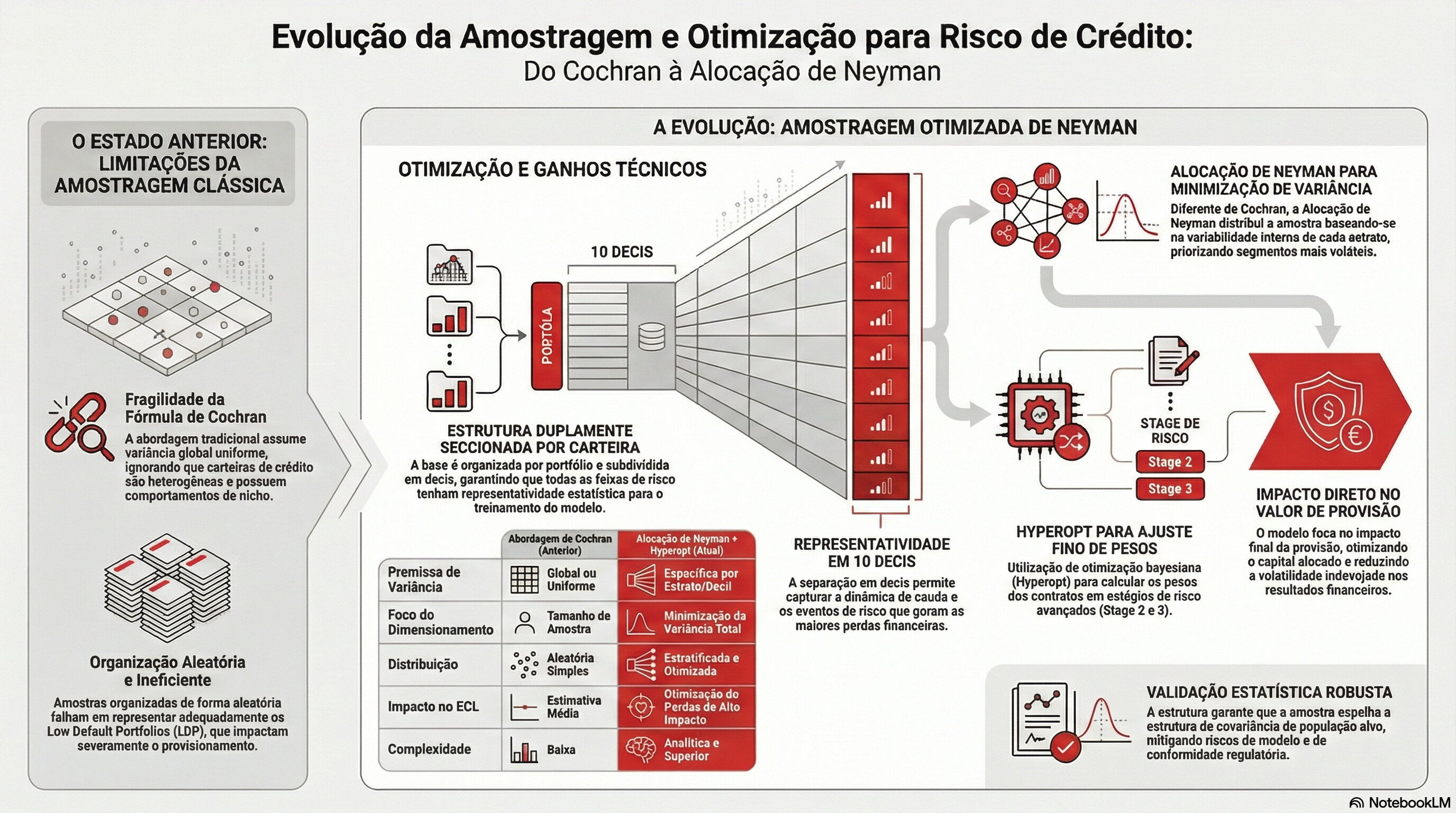
Mathematical Foundation:
Cochran (1977) infinite-population formula:
n_0 = (Z² × σ²) / E²
Finite-population correction (FPC):
n = n_0 / (1 + (n_0 - 1) / N)
Stratified Neyman-adjusted minimum:
n_strat = Σ_h [ (N_h × S_h)² / n_0_h ] / (Σ N_h × S_h)²
then corrected for FPC per stratum.
Where:
Z = Z-score for the desired confidence level
σ² = population variance of the target variable
E = margin of error (relative to the population mean)
N = total population size
N_h = population of stratum h (per stage)
S_h = standard deviation in stratum h
Args:
df (DataFrame): Full filtered population DataFrame.
target_col (str): Column to compute variance on.
stage_col (str): Column defining strata for stratified calc.
confidence_level (float): Desired confidence (e.g., 0.95 → Z=1.96).
margin_of_error (float): Desired relative precision (0.05 = 5%).
Expected Input Schema:
DataFrame with numeric ``target_col`` and categorical ``stage_col``.
Output Schema Transformation:
Returns a dict with:
- ``n_cochran_infinite`` (int): Cochran infinite-pop minimum.
- ``n_cochran_fpc`` (int): Cochran with FPC.
- ``n_stratified_neyman`` (int): Stratified Neyman minimum.
- ``population_N`` (int): Total population count.
- ``population_mean`` (float): Population mean of target.
- ``population_stddev`` (float): Population stddev of target.
- ``per_stage`` (Dict): Per-stage breakdown with N_h, S_h, n_h.
Memory Implications:
Two aggregations (global + per-stage). Output is ~4 rows on
the driver. Negligible.
Exception Handling Protocols:
Returns conservative estimates (n=N) if variance is zero or
mean is zero (avoiding division by zero).
Returns:
Dict[str, Any]: Minimum sample size report.
"""
# ── Z-score lookup for common confidence levels ──
z_map = {0.90: 1.645, 0.95: 1.960, 0.99: 2.576}
z_score = z_map.get(confidence_level, 1.960)
# ── Global population statistics (100% PySpark) ──
global_stats = df.agg(
F.count(target_col).alias("N"),
F.mean(target_col).alias("mu"),
F.stddev(target_col).alias("sigma"),
F.variance(target_col).alias("var"),
).collect()[0]
N = global_stats["N"]
mu = global_stats["mu"] if global_stats["mu"] is not None else 1.0
sigma = global_stats["sigma"] if global_stats["sigma"] is not None else 0.0
var = global_stats["var"] if global_stats["var"] is not None else 0.0
if N == 0 or mu == 0 or sigma == 0:
print(" ⚠ Variância ou média zero — retornando N como mínimo.")
return {
"n_cochran_infinite": N,
"n_cochran_fpc": N,
"n_stratified_neyman": N,
"population_N": N,
"population_mean": float(mu),
"population_stddev": float(sigma),
"per_stage": {},
}
# Absolute margin of error = relative margin × mean
E_abs = margin_of_error * abs(mu)
# ── Cochran Infinite Population ──
n_0 = math.ceil((z_score ** 2 * var) / (E_abs ** 2))
# ── Cochran with Finite Population Correction ──
n_fpc = math.ceil(n_0 / (1.0 + (n_0 - 1.0) / N))
# ── Per-Stage Statistics (Stratified) ──
stage_stats = df.groupBy(stage_col).agg(
F.count(target_col).alias("N_h"),
F.stddev(target_col).alias("S_h"),
).collect()
per_stage: Dict[str, Dict[str, Any]] = {}
sum_nh_sh = 0.0
sum_nh_sh2 = 0.0
for row in stage_stats:
stage_val = str(row[stage_col])
nh = row["N_h"]
sh = row["S_h"] if row["S_h"] is not None and row["S_h"] > 0 else 1e-9
sum_nh_sh += nh * sh
sum_nh_sh2 += nh * (sh ** 2)
per_stage[stage_val] = {"N_h": nh, "S_h": round(sh, 4)}
# ── Stratified Neyman Minimum ──
# n_neyman = (Σ N_h × S_h)² / (N² × D + Σ N_h × S_h²)
# where D = E² / Z²
D = (E_abs ** 2) / (z_score ** 2)
n_neyman_denom = (N ** 2) * D + sum_nh_sh2
n_neyman = math.ceil((sum_nh_sh ** 2) / n_neyman_denom) if n_neyman_denom > 0 else N
# Per-stage quota (proportional to N_h × S_h)
for stage_val, info in per_stage.items():
nh = info["N_h"]
sh = info["S_h"]
if sum_nh_sh > 0:
quota = math.ceil(n_neyman * (nh * sh) / sum_nh_sh)
else:
quota = math.ceil(n_neyman / len(per_stage))
per_stage[stage_val]["n_h_minimum"] = min(quota, nh)
result = {
"n_cochran_infinite": n_0,
"n_cochran_fpc": n_fpc,
"n_stratified_neyman": n_neyman,
"population_N": N,
"population_mean": round(float(mu), 4),
"population_stddev": round(float(sigma), 4),
"per_stage": per_stage,
}
# ── Print Summary ──
print(f"\n ┌── TAMANHO MÍNIMO AMOSTRAL (Cochran + Neyman Estratificado)")
print(f" │ População N = {N:>12,}")
print(f" │ Média (μ) = {mu:>12,.2f}")
print(f" │ Desvio Padrão (σ) = {sigma:>12,.2f}")
print(f" │ Confiança = {confidence_level*100:.0f}% (Z={z_score})")
print(f" │ Margem de Erro = {margin_of_error*100:.1f}% (E_abs={E_abs:,.2f})")
print(f" │ ────────────────────────────────────────")
print(f" │ n₀ (Cochran ∞) = {n_0:>12,}")
print(f" │ n (Cochran FPC) = {n_fpc:>12,}")
print(f" │ n (Neyman Estrat.) = {n_neyman:>12,}")
print(f" │ ────────────────────────────────────────")
for stage_val in sorted(per_stage.keys()):
info = per_stage[stage_val]
print(
f" │ Stage {stage_val:>3s}: "
f"N_h={info['N_h']:>8,} S_h={info['S_h']:>10.4f} "
f"n_h_min={info['n_h_minimum']:>6,}"
)
print(f" └────────────────────────────────────────────────")
return result
# ╔═════════════════════════════════════════════════════════════════════════╗
# ║ 2B. DATA QUALITY FEATURE PROFILER (CHUNKED, OOM-SAFE) ║
# ╚═════════════════════════════════════════════════════════════════════════╝
class DataQualityProfiler:
"""
Distributed Data Quality & Feature Profiler for wide IFRS 9 tables.
Processes 500+ columns safely by splitting them into configurable
chunks (default: 50 columns per iteration) to prevent OOM on both
the Driver and Executors. Each chunk produces a tall/melted DataFrame
of per-column metrics, which are unioned into a single profiling table.
Metrics Computed:
**Numeric columns** (types: double, int, float, long, decimal):
- ``missing_pct``: Percentage of null/NaN values.
- ``zero_pct``: Percentage of exact-zero values.
- ``min_val``, ``max_val``, ``mean_val``: Basic statistics.
- ``q25``, ``q50_median``, ``q75``: Approximate quantiles
via ``approxQuantile`` (T-Digest, ε=0.01).
- ``iqr``: Interquartile Range (Q75 - Q25).
**Categorical columns** (types: string, boolean):
- ``missing_pct``: Percentage of null values.
- ``distinct_count``: Count of distinct non-null values.
- ``top_n_values``: Top-N most frequent values as a
comma-separated string (100% PySpark, no toPandas).
OOM Prevention Strategy:
With 500 columns, a single ``df.agg(...)`` call would create a
Catalyst logical plan with ~3000 expressions (6 per column),
which overwhelms the JVM's code generation and the Driver's
memory. By chunking into groups of 50, each aggregation has
~300 expressions — well within safe limits. The results are
unioned as tall rows (column_name, metric_name, metric_value),
keeping the Driver payload to ~50KB per chunk.
Attributes:
spark (SparkSession): Active session.
config (SamplerConfig): Suite configuration.
"""
# ── Numeric Spark types (checked via string prefix) ──
_NUMERIC_TYPE_PREFIXES = ("double", "float", "int", "long", "bigint",
"short", "decimal", "tinyint", "smallint")
def __init__(self, spark: SparkSession, config: SamplerConfig) -> None:
"""Initializes the profiler.
Args:
spark (SparkSession): Active Spark session.
config (SamplerConfig): Suite configuration.
Expected Input Schema:
None. Configuration only.
Output Schema Transformation:
None.
Memory Implications:
Negligible.
Exception Handling Protocols:
None.
"""
self.spark = spark
self.config = config
def _classify_columns(
self, df: DataFrame,
) -> Tuple[List[str], List[str]]:
"""Splits DataFrame columns into numeric and categorical lists.
Args:
df (DataFrame): Source DataFrame to classify.
Expected Input Schema:
Any DataFrame.
Output Schema Transformation:
Returns two lists: (numeric_cols, categorical_cols).
Memory Implications:
Iterates schema metadata on Driver. Negligible.
Exception Handling Protocols:
Columns of type ``date``, ``timestamp``, ``binary``, ``array``,
``map``, ``struct`` are excluded from both lists.
Returns:
Tuple[List[str], List[str]]: (numeric, categorical) columns.
"""
numeric_cols: List[str] = []
categorical_cols: List[str] = []
exclude_types = ("date", "timestamp", "binary", "array", "map", "struct")
for field in df.schema.fields:
type_str = field.dataType.simpleString().lower()
if any(type_str.startswith(ex) for ex in exclude_types):
continue
if any(type_str.startswith(nt) for nt in self._NUMERIC_TYPE_PREFIXES):
numeric_cols.append(field.name)
elif type_str in ("string", "boolean"):
categorical_cols.append(field.name)
return numeric_cols, categorical_cols
def _profile_numeric_chunk(
self, df: DataFrame, cols: List[str], stage_val: int, total_count: int,
) -> DataFrame:
"""Profiles a chunk of numeric columns for a single stage.
Args:
df (DataFrame): Stage-filtered DataFrame.
cols (List[str]): Column names in this chunk.
stage_val (int): Current IFRS stage value.
total_count (int): Pre-computed row count for the stage.
Expected Input Schema:
DataFrame with all columns in ``cols`` as numeric types.
Output Schema Transformation:
Returns a tall DataFrame with columns:
``[carteira, stage, column_name, col_type, missing_pct,
zero_pct, min_val, max_val, mean_val, q25, q50_median,
q75, iqr, distinct_count, top_n_values]``.
Memory Implications:
One ``agg()`` call with ~6 expressions per column. With
chunk_size=50, that's ~300 expressions — safe for Catalyst.
Exception Handling Protocols:
Columns that fail aggregation are logged and skipped.
Returns:
DataFrame: Tall profiling rows for the chunk.
"""
cart = self.config.carteira_ifrs
results: List[Dict[str, Any]] = []
# ── Build aggregation expressions in one pass ──
agg_exprs = []
for c in cols:
agg_exprs.extend([
F.sum(F.when(F.col(c).isNull(), 1).otherwise(0)).alias(f"{c}__null_cnt"),
F.sum(F.when(F.col(c) == 0, 1).otherwise(0)).alias(f"{c}__zero_cnt"),
F.min(c).alias(f"{c}__min"),
F.max(c).alias(f"{c}__max"),
F.mean(c).alias(f"{c}__mean"),
])
stats_row = df.agg(*agg_exprs).collect()[0]
# ── Quantiles via approxQuantile (one call per column) ──
for c in cols:
try:
quantiles = df.approxQuantile(c, [0.25, 0.50, 0.75], 0.01)
q25 = quantiles[0] if len(quantiles) > 0 else None
q50 = quantiles[1] if len(quantiles) > 1 else None
q75 = quantiles[2] if len(quantiles) > 2 else None
except Exception:
q25, q50, q75 = None, None, None
null_cnt = stats_row[f"{c}__null_cnt"] or 0
zero_cnt = stats_row[f"{c}__zero_cnt"] or 0
results.append({
"carteira": cart,
"stage": stage_val,
"column_name": c,
"col_type": "numeric",
"missing_pct": round(null_cnt / total_count, 6) if total_count > 0 else 0.0,
"zero_pct": round(zero_cnt / total_count, 6) if total_count > 0 else 0.0,
"min_val": float(stats_row[f"{c}__min"]) if stats_row[f"{c}__min"] is not None else None,
"max_val": float(stats_row[f"{c}__max"]) if stats_row[f"{c}__max"] is not None else None,
"mean_val": float(stats_row[f"{c}__mean"]) if stats_row[f"{c}__mean"] is not None else None,
"q25": float(q25) if q25 is not None else None,
"q50_median": float(q50) if q50 is not None else None,
"q75": float(q75) if q75 is not None else None,
"iqr": float(q75 - q25) if q25 is not None and q75 is not None else None,
"distinct_count": None,
"top_n_values": None,
})
return self.spark.createDataFrame(results)
def _profile_categorical_chunk(
self, df: DataFrame, cols: List[str], stage_val: int, total_count: int,
) -> DataFrame:
"""Profiles a chunk of categorical columns for a single stage.
Extracts Top-N values using a distributed groupBy + Window rank
strategy: for each column, count values via groupBy, rank by
frequency, and collect the top N as a comma-separated string.
This avoids ``collect_list`` on high-cardinality columns which
would OOM the Driver.
Args:
df (DataFrame): Stage-filtered DataFrame.
cols (List[str]): Column names in this chunk.
stage_val (int): Current IFRS stage value.
total_count (int): Pre-computed row count for the stage.
Expected Input Schema:
DataFrame with all columns in ``cols`` as string/boolean.
Output Schema Transformation:
Returns a tall DataFrame with same schema as numeric chunk.
Memory Implications:
One ``agg()`` for null counts + distinct counts (~2 exprs
per column). One groupBy per column for top-N (bounded by
``dq_top_n_categorical`` rows collected per column).
Exception Handling Protocols:
Errors in top-N extraction are caught; ``top_n_values``
defaults to "ERROR".
Returns:
DataFrame: Tall profiling rows for the chunk.
"""
cart = self.config.carteira_ifrs
top_n = self.config.dq_top_n_categorical
results: List[Dict[str, Any]] = []
# ── Aggregation: null count + distinct count ──
agg_exprs = []
for c in cols:
agg_exprs.extend([
F.sum(F.when(F.col(c).isNull(), 1).otherwise(0)).alias(f"{c}__null_cnt"),
F.countDistinct(c).alias(f"{c}__dist"),
])
stats_row = df.agg(*agg_exprs).collect()[0]
# ── Top-N extraction per column (bounded collect) ──
for c in cols:
null_cnt = stats_row[f"{c}__null_cnt"] or 0
dist_cnt = stats_row[f"{c}__dist"] or 0
# Top-N: groupBy → count → orderBy desc → limit → collect
top_n_str = ""
try:
top_rows = (
df.filter(F.col(c).isNotNull())
.groupBy(c)
.agg(F.count("*").alias("_freq"))
.orderBy(F.desc("_freq"))
.limit(top_n)
.collect()
)
top_n_str = ", ".join(
f"{row[c]}({row['_freq']:,})" for row in top_rows
)
except Exception as e:
top_n_str = f"ERROR: {e}"
results.append({
"carteira": cart,
"stage": stage_val,
"column_name": c,
"col_type": "categorical",
"missing_pct": round(null_cnt / total_count, 6) if total_count > 0 else 0.0,
"zero_pct": None,
"min_val": None,
"max_val": None,
"mean_val": None,
"q25": None,
"q50_median": None,
"q75": None,
"iqr": None,
"distinct_count": int(dist_cnt),
"top_n_values": top_n_str,
})
return self.spark.createDataFrame(results)
def run_profiling(
self, df: DataFrame,
) -> DataFrame:
"""Executes the full Data Quality profiling pipeline.
Iterates over all IFRS Stages present in the data, and for
each stage processes columns in chunks of ``dq_chunk_size``
to avoid OOM.
Args:
df (DataFrame): Full filtered/cast population DataFrame
(already filtered by carteira and date).
Expected Input Schema:
Any DataFrame with a ``stage`` column.
Output Schema Transformation:
Returns a tall DataFrame with one row per (stage, column)
containing all quality metrics.
Memory Implications:
Processes ``ceil(n_cols / chunk_size)`` aggregations per
stage. With 500 cols and chunk=50, that's 10 iterations
per stage × 3 stages = 30 total aggregation jobs.
Each job touches the full partition for that stage but
only computes ~300 Catalyst expressions.
Exception Handling Protocols:
Stage-level failures are caught and logged. Partial
results from successful stages are preserved.
Returns:
DataFrame: Complete profiling results.
"""
chunk_size = self.config.dq_chunk_size
stage_col = self.config.stage_col
cart = self.config.carteira_ifrs
print(f"\n{'='*72}")
print(f" DATA QUALITY PROFILER — Carteira: {cart}")
print(f" Chunk Size: {chunk_size} colunas por iteração")
print(f"{'='*72}")
# ── Classify all columns ──
numeric_cols, categorical_cols = self._classify_columns(df)
total_cols = len(numeric_cols) + len(categorical_cols)
print(f" ├── Colunas detectadas: {total_cols} total")
print(f" │ Numéricas: {len(numeric_cols)}")
print(f" │ Categóricas: {len(categorical_cols)}")
# ── Get distinct stages ──
stages = sorted(
[row[stage_col] for row in df.select(stage_col).distinct().collect()]
)
print(f" ├── Stages encontrados: {stages}")
all_chunks: List[DataFrame] = []
for stg in stages:
print(f"\n ┌── Profiling Stage {stg}")
df_stage = df.filter(F.col(stage_col) == stg)
df_stage.cache()
stage_count = df_stage.count()
print(f" │ Registros: {stage_count:,}")
# ── Numeric chunks ──
n_num_chunks = math.ceil(len(numeric_cols) / chunk_size) if numeric_cols else 0
for i in range(n_num_chunks):
chunk = numeric_cols[i * chunk_size: (i + 1) * chunk_size]
print(f" │ Chunk numérico {i+1}/{n_num_chunks}: {len(chunk)} colunas")
try:
chunk_df = self._profile_numeric_chunk(
df_stage, chunk, stg, stage_count,
)
all_chunks.append(chunk_df)
except Exception as e:
print(f" │ ✗ Falha no chunk numérico {i+1}: {e}")
# ── Categorical chunks ──
n_cat_chunks = math.ceil(len(categorical_cols) / chunk_size) if categorical_cols else 0
for i in range(n_cat_chunks):
chunk = categorical_cols[i * chunk_size: (i + 1) * chunk_size]
print(f" │ Chunk categórico {i+1}/{n_cat_chunks}: {len(chunk)} colunas")
try:
chunk_df = self._profile_categorical_chunk(
df_stage, chunk, stg, stage_count,
)
all_chunks.append(chunk_df)
except Exception as e:
print(f" │ ✗ Falha no chunk categórico {i+1}: {e}")
df_stage.unpersist()
print(f" └── Stage {stg} concluído.")
if not all_chunks:
print(" ⚠ Nenhum resultado de profiling gerado.")
return self.spark.createDataFrame([], StructType([]))
# ── Union all chunks ──
result_df = all_chunks[0]
for chunk_df in all_chunks[1:]:
result_df = result_df.unionByName(chunk_df, allowMissingColumns=True)
# ── Persist to Delta ──
date_suffix = datetime.now().strftime("%Y%m")
cart_safe = cart.lower().replace(" ", "_")
table_name = f"tb_dq_profiling_{cart_safe}_{date_suffix}"
full_table = f"{self.config.dest_schema}.{table_name}"
result_df.write.format("delta").mode("overwrite").option(
"overwriteSchema", "true",
).saveAsTable(full_table)
n_rows = result_df.count()
print(f"\n ✔ Profiling salvo: {full_table} → {n_rows:,} métricas")
return result_df
def get_usable_columns(
self, profiling_df: DataFrame, stage_val: int,
max_missing_pct: Optional[float] = None,
) -> List[str]:
"""Extracts the list of columns passing the missing threshold.
Given a profiling results DataFrame, filters to the requested
stage and returns only column names with ``missing_pct`` below
the configured threshold.
Args:
profiling_df (DataFrame): Output from ``run_profiling()``.
stage_val (int): Stage to filter.
max_missing_pct (Optional[float]): Override threshold.
If None, uses ``config.dq_missing_threshold``.
Expected Input Schema:
DataFrame with ``stage``, ``column_name``, ``missing_pct``.
Output Schema Transformation:
Returns a plain list of column name strings.
Memory Implications:
Collects ~500 rows max (one per column per stage). ~20KB.
Exception Handling Protocols:
Returns empty list if profiling_df is empty.
Returns:
List[str]: Column names that pass the quality gate.
"""
threshold = max_missing_pct if max_missing_pct is not None \
else self.config.dq_missing_threshold
usable_rows = (
profiling_df
.filter(
(F.col("stage") == stage_val)
& (F.col("missing_pct") <= threshold)
)
.select("column_name")
.distinct()
.collect()
)
usable_cols = [row["column_name"] for row in usable_rows]
print(
f" │ Stage {stage_val}: {len(usable_cols)} colunas com "
f"missing ≤ {threshold*100:.0f}%"
)
return usable_cols
# ╔═════════════════════════════════════════════════════════════════════════╗
# ║ 2C. COLUMN TYPE ANALYZER — AUTO-DETECTION ENGINE ║
# ╚═════════════════════════════════════════════════════════════════════════╝
@dataclass
class ColumnClassification:
"""Immutable container for automatic column type classification.
Attributes:
numeric (List[str]): Native numeric columns (int, double, float,
long, decimal, short, tinyint, smallint, bigint).
numeric_inferred (List[str]): String columns whose non-null
values successfully cast to ``double`` above the configured
threshold (default 95%). These were stored as ``string``
in Delta but are semantically numeric.
flag_binary (List[str]): String columns with ≤ ``flag_max_distinct``
unique non-null values, all matching known flag patterns
(e.g., "0"/"1", "S"/"N", "true"/"false").
categorical (List[str]): Plain string/boolean columns that do
NOT contain the ``#`` vectorised pattern and are not numeric
nor flag.
numeric_vectorized (List[str]): String columns whose non-null
values are ``#``-separated numeric elements (e.g.,
``"0.01#0.02#0.03"``).
categorical_vectorized (List[str]): String columns whose non-null
values are ``#``-separated textual elements (e.g.,
``"AAA#AA#A"``).
temporal (List[str]): Date and timestamp columns.
date_inferred (List[str]): String columns whose values parse
into valid dates using configured format patterns.
excluded (List[str]): Complex types (array, map, struct, binary)
that cannot be profiled or sampled.
vector_lengths (Dict[str, int]): Maximum element count discovered
per vectorised column. Used by ``VectorizedColumnHandler`` to
determine how many sub-columns to create.
sample_values (Dict[str, List[str]]): Up to N sample non-null
values per column for visual validation by the user.
Memory Implications:
Negligible. Holds only column name lists and small dicts.
Exception Handling Protocols:
None. Pure data container.
"""
numeric: List[str] = field(default_factory=list)
numeric_inferred: List[str] = field(default_factory=list)
flag_binary: List[str] = field(default_factory=list)
categorical: List[str] = field(default_factory=list)
numeric_vectorized: List[str] = field(default_factory=list)
categorical_vectorized: List[str] = field(default_factory=list)
temporal: List[str] = field(default_factory=list)
date_inferred: List[str] = field(default_factory=list)
excluded: List[str] = field(default_factory=list)
vector_lengths: Dict[str, int] = field(default_factory=dict)
sample_values: Dict[str, List[str]] = field(default_factory=dict)
class ColumnTypeAnalyzer:
"""Distributed Column Type Detection Engine.
Executes a 3-phase analysis pipeline to classify every column in a
DataFrame into one of nine semantic categories.
Phase 1 — Schema Inspection (O(1), Driver-only):
Classifies columns by native Spark data types. Numeric types
(int, double, float, long, decimal …) are labelled immediately.
String columns are forwarded to Phase 2 for deep analysis.
Phase 2 — Deep String / Vectorised Analysis (Distributed, Chunked):
For each chunk of ``vectorized_chunk_size`` string columns,
computes **4 metrics in a single distributed ``agg()``**:
1. ``nonnull_count``: Non-null row count.
2. ``separator_count``: Rows containing the ``#`` separator.
3. ``max_vector_length``: ``max(size(split(col, '#')))``.
4. ``numeric_first_element``: Rows where the first split
element successfully casts to ``double``.
Classification Rules:
- ``sep_count / nonnull >= vectorized_min_pct`` (default 50%)
AND ``max_vector_length > 1`` → **vectorized**.
- Of vectorized: ``numeric_first / sep_count >=
vectorized_numeric_threshold`` (default 90%)
→ **numeric_vectorized**.
- Otherwise → **categorical_vectorized**.
- Non-vectorized strings are forwarded to Phase 3.
Phase 3 — Semantic Re-Typing (Distributed, Chunked):
For each non-vectorised string column, computes **5 metrics**
in a single distributed ``agg()``:
1. ``nonnull_count``: Non-null row count.
2. ``castable_count``: Rows where ``cast(col as double)``
succeeds (i.e. result is not null).
3. ``approx_distinct``: Approximate count of distinct non-null
values (via ``approx_count_distinct``).
4. ``trimmed_lower_set``: Collected distinct ``lower(trim(col))``
values (only for columns with ≤ ``flag_max_distinct``).
5. ``date_castable_count``: For columns failing numeric, tries
``to_date`` with configured format patterns.
Classification Rules:
- ``castable / nonnull >= string_to_numeric_threshold``
(default 95%) → **numeric_inferred**.
- ``approx_distinct <= flag_max_distinct`` AND all values
match ``flag_known_values`` → **flag_binary**.
- Any format in ``date_sample_formats`` parses ≥ 95% of
rows → **date_inferred**.
- Otherwise → **categorical**.
OOM Prevention:
All phases use chunked aggregation to keep Catalyst expression
count per ``agg()`` within safe limits.
Attributes:
spark (SparkSession): Active session.
config (SamplerConfig): Suite configuration.
"""
_NUMERIC_PREFIXES = (
"double", "float", "int", "long", "bigint",
"short", "decimal", "tinyint", "smallint",
)
_TEMPORAL_PREFIXES = ("date", "timestamp")
_COMPLEX_PREFIXES = ("array", "map", "struct", "binary")
def __init__(self, spark: SparkSession, config: SamplerConfig) -> None:
"""Initializes the analyzer.
Args:
spark (SparkSession): Active Spark session.
config (SamplerConfig): Suite configuration.
Expected Input Schema:
None. Configuration only.
Output Schema Transformation:
None.
Memory Implications:
Negligible. Stores references.
Exception Handling Protocols:
None.
"""
self.spark = spark
self.config = config
def analyze(self, df: DataFrame) -> ColumnClassification:
"""Execute the full column classification pipeline.
Args:
df (DataFrame): Population DataFrame (post-filter).
Expected Input Schema:
Any DataFrame.
Output Schema Transformation:
Returns ``ColumnClassification``.
Persists to Delta:
``{dest_schema}.tb_column_classification_{cart}_{YYYYMM}``.
Memory Implications:
Phase 1: O(1) on Driver.
Phase 2: ``ceil(n_string / chunk_size)`` distributed aggs,
each with 4 expressions per column.
Exception Handling Protocols:
Chunk-level failures are caught; columns in failed chunks
default to ``categorical``.
Returns:
ColumnClassification: Full classification result.
"""
total_cols = len(df.columns)
print(f"\n{'=' * 72}")
print(f" COLUMN TYPE ANALYZER — Detecção Automática ({total_cols} Colunas)")
print(f"{'=' * 72}")
# ── Phase 1: Schema Classification ──
print(f"\n ┌── Fase 1: Classificação por Schema Spark")
schema_map = self._phase1_schema_classify(df)
print(f" │ Numéricas Nativas: {len(schema_map['numeric']):>4}")
print(f" │ String (candidatas): {len(schema_map['string']):>4}")
print(f" │ Boolean: {len(schema_map['boolean']):>4}")
print(f" │ Temporal: {len(schema_map['temporal']):>4}")
print(f" │ Complexas/Excl.: {len(schema_map['complex']):>4}")
# ── Process Manual Hints ──
man_num = set(self.config.manual_numeric_cols)
man_cat = set(self.config.manual_categorical_cols)
man_flag = set(self.config.manual_flag_cols)
man_vnum = set(self.config.manual_vectorized_numeric_cols)
man_vcat = set(self.config.manual_vectorized_categorical_cols)
man_date = set(self.config.manual_date_cols)
all_man = man_num | man_cat | man_flag | man_vnum | man_vcat | man_date
# Remove manual columns from strings to avoid deep processing
strings_to_process = [c for c in schema_map["string"] if c not in all_man]
# ── Phase 2: Deep String / Vectorised Analysis ──
vec_num: List[str] = []
vec_cat: List[str] = []
non_vec_strings: List[str] = []
vec_lengths: Dict[str, int] = {}
if strings_to_process:
n_str = len(strings_to_process)
sep = self.config.vectorized_separator
print(f"\n ├── Fase 2: Análise Profunda de {n_str} Colunas String")
print(
f" │ Separador: '{sep}' | "
f"Limiar Vec.: {self.config.vectorized_min_pct * 100:.0f}% | "
f"Limiar Num.: {self.config.vectorized_numeric_threshold * 100:.0f}%"
)
vec_num, vec_cat, non_vec_strings, vec_lengths = \
self._phase2_deep_string_analysis(df, strings_to_process)
# ── Phase 3: Semantic Re-Typing (non-vectorised strings) ──
num_inferred: List[str] = []
flag_binary: List[str] = []
date_inferred: List[str] = []
pure_cat: List[str] = []
all_non_vec = non_vec_strings + schema_map["boolean"]
if all_non_vec:
print(
f"\n ├── Fase 3: Retipagem Semântica de "
f"{len(all_non_vec)} Colunas Não-Vetorizadas"
)
print(
f" │ Limiar Num.: "
f"{self.config.string_to_numeric_threshold * 100:.0f}% | "
f"Max Distinct Flag: {self.config.flag_max_distinct}"
)
num_inferred, flag_binary, date_inferred, pure_cat = \
self._phase3_semantic_retyping(df, all_non_vec)
# ── Phase 4: Collect Sample Values (only for key types) ──
sample_values: Dict[str, List[str]] = {}
if self.config.enable_sample_collection:
print(f"\n ├── Fase 4: Coleta de Valores de Exemplo")
# Only collect samples for types that need visual validation
# Skip native numeric/temporal (those are obviously correct)
sample_candidates: Dict[str, str] = {}
for c in num_inferred:
sample_candidates[c] = "numeric_inferred"
for c in flag_binary:
sample_candidates[c] = "flag_binary"
for c in vec_num[:10]: # limit vector samples
sample_candidates[c] = "numeric_vectorized"
for c in vec_cat[:10]:
sample_candidates[c] = "categorical_vectorized"
for c in date_inferred:
sample_candidates[c] = "date_inferred"
for c in pure_cat[:20]: # only first 20 categoricals
sample_candidates[c] = "categorical"
sample_values = self._collect_sample_values(
df, sample_candidates,
)
else:
print(f"\n ├── Fase 4: Coleta de exemplos desabilitada")
# ── Add Manual Hints back into final arrays ──
num_inferred.extend([c for c in man_num if c in schema_map["string"]])
pure_cat.extend([c for c in man_cat if c in schema_map["string"]])
flag_binary.extend([c for c in man_flag if c in schema_map["string"]])
vec_num.extend([c for c in man_vnum if c in schema_map["string"]])
vec_cat.extend([c for c in man_vcat if c in schema_map["string"]])
date_inferred.extend([c for c in man_date if c in schema_map["string"]])
# ── Build Result ──
clf = ColumnClassification(
numeric=schema_map["numeric"],
numeric_inferred=num_inferred,
flag_binary=flag_binary,
categorical=pure_cat,
numeric_vectorized=vec_num,
categorical_vectorized=vec_cat,
temporal=schema_map["temporal"],
date_inferred=date_inferred,
excluded=schema_map["complex"],
vector_lengths=vec_lengths,
sample_values=sample_values,
)
self._print_classification_report(clf, total_cols)
self._print_detailed_sample_report(clf)
self._persist_classification(clf)
return clf
# ── Internal Methods ──────────────────────────────────────────────
def _phase1_schema_classify(
self, df: DataFrame,
) -> Dict[str, List[str]]:
"""Classifies columns purely by native Spark schema type.
Args:
df (DataFrame): Input DataFrame.
Expected Input Schema:
Any DataFrame.
Output Schema Transformation:
Returns dict with five keys: ``numeric``, ``string``,
``boolean``, ``temporal``, ``complex``.
Memory Implications:
Iterates ``df.schema.fields`` on Driver. O(n_cols). Negligible.
Exception Handling Protocols:
Unknown types are placed in ``complex``.
Returns:
Dict[str, List[str]]: Schema-based classification.
"""
result: Dict[str, List[str]] = {
"numeric": [], "string": [], "boolean": [],
"temporal": [], "complex": [],
}
for fld in df.schema.fields:
ts = fld.dataType.simpleString().lower()
if any(ts.startswith(p) for p in self._NUMERIC_PREFIXES):
result["numeric"].append(fld.name)
elif ts == "string":
result["string"].append(fld.name)
elif ts == "boolean":
result["boolean"].append(fld.name)
elif any(ts.startswith(p) for p in self._TEMPORAL_PREFIXES):
result["temporal"].append(fld.name)
else:
result["complex"].append(fld.name)
return result
def _phase2_deep_string_analysis(
self, df: DataFrame, string_cols: List[str],
) -> Tuple[List[str], List[str], List[str], Dict[str, int]]:
"""Detects vectorised patterns in string columns via distributed agg.
For each chunk, computes 4 metrics in ONE distributed aggregation:
(1) non-null count
(2) separator presence count
(3) max split size (vector length)
(4) numeric first-element cast success count
Args:
df (DataFrame): Input DataFrame.
string_cols (List[str]): String columns from Phase 1.
Expected Input Schema:
DataFrame with all ``string_cols`` as ``StringType``.
Output Schema Transformation:
Returns 4-tuple:
``(vec_numeric, vec_categorical, pure_categorical,
vector_lengths_dict)``.
Memory Implications:
``ceil(n_cols / chunk)`` aggregations. With chunk=30
→ ~120 Catalyst expressions per agg. Safe.
Exception Handling Protocols:
Failed chunks default all member columns to categorical.
Returns:
Tuple: Classification lists and vector length dict.
"""
sep = self.config.vectorized_separator
escaped_sep = re.escape(sep)
chunk_size = self.config.vectorized_chunk_size
min_pct = self.config.vectorized_min_pct
num_threshold = self.config.vectorized_numeric_threshold
vec_numeric: List[str] = []
vec_categorical: List[str] = []
non_vectorized: List[str] = []
vec_lengths: Dict[str, int] = {}
n_chunks = math.ceil(len(string_cols) / chunk_size)
for ci in range(n_chunks):
chunk = string_cols[ci * chunk_size: (ci + 1) * chunk_size]
print(
f" │ Chunk {ci + 1}/{n_chunks}: "
f"{len(chunk)} colunas..."
)
# Track column-name → sanitised alias key
col_keys: List[Tuple[str, str]] = []
try:
agg_exprs = []
for c in chunk:
key = (
c.replace(".", "__")
.replace(" ", "_")
.replace("-", "_")
)
col_keys.append((c, key))
cr = F.col(f"`{c}`")
# (1) Non-null count
agg_exprs.append(
F.sum(
F.when(cr.isNotNull(), 1).otherwise(0)
).alias(f"{key}___nn")
)
# (2) Separator presence count
agg_exprs.append(
F.sum(F.when(
cr.isNotNull() & (F.instr(cr, sep) > 0), 1,
).otherwise(0)).alias(f"{key}___sc")
)
# (3) Max vector length
agg_exprs.append(
F.max(F.when(
F.instr(cr, sep) > 0,
F.size(F.split(cr, escaped_sep)),
)).alias(f"{key}___ml")
)
# (4) Numeric first element (cast test)
agg_exprs.append(
F.sum(F.when(
(F.instr(cr, sep) > 0)
& F.element_at(
F.split(cr, escaped_sep), 1,
).cast("double").isNotNull(),
1,
).otherwise(0)).alias(f"{key}___nf")
)
# Single distributed aggregation pass
stats = df.agg(*agg_exprs).collect()[0]
for c, key in col_keys:
nn = stats[f"{key}___nn"] or 0
sc = stats[f"{key}___sc"] or 0
ml = stats[f"{key}___ml"]
nf = stats[f"{key}___nf"] or 0
if nn == 0:
non_vectorized.append(c)
continue
sep_ratio = sc / nn
if sep_ratio >= min_pct and ml is not None and ml > 1:
vec_lengths[c] = int(ml)
num_ratio = nf / sc if sc > 0 else 0.0
if num_ratio >= num_threshold:
vec_numeric.append(c)
print(
f" │ ✔ {c}: NUMÉRICO VETORIZADO "
f"(len={ml}, vec={sep_ratio:.0%}, "
f"num={num_ratio:.0%})"
)
else:
vec_categorical.append(c)
print(
f" │ ✔ {c}: CATEGÓRICO VETORIZADO "
f"(len={ml}, vec={sep_ratio:.0%})"
)
else:
non_vectorized.append(c)
except Exception as e:
print(f" │ ✗ Chunk {ci + 1} falhou: {e}")
for c, _ in col_keys:
if c not in non_vectorized:
non_vectorized.append(c)
return vec_numeric, vec_categorical, non_vectorized, vec_lengths
def _phase3_semantic_retyping(
self, df: DataFrame, non_vec_cols: List[str],
) -> Tuple[List[str], List[str], List[str], List[str]]:
"""Semantic re-typing of non-vectorised string columns.
For each chunk, computes 3 core metrics in ONE distributed agg:
(1) non-null count
(2) castable-to-double count (whole value, not split)
(3) approx_count_distinct
Columns that fail numeric detection are tested for flag/boolean
patterns via a second lightweight aggregation that collects
distinct values (only for low-cardinality columns).
Columns that fail both are tested for date parsability.
Args:
df (DataFrame): Input DataFrame.
non_vec_cols (List[str]): String columns from Phase 2 that
are NOT vectorised.
Expected Input Schema:
DataFrame with all ``non_vec_cols`` as ``StringType``.
Output Schema Transformation:
Returns 4-tuple:
``(numeric_inferred, flag_binary, date_inferred,
pure_categorical)``.
Memory Implications:
``ceil(n_cols / chunk_size)`` distributed aggs, each with
3 expressions per column. Safe for 500+ columns.
Exception Handling Protocols:
Chunk-level failures default columns to categorical.
Returns:
Tuple: Classification lists.
"""
chunk_size = self.config.vectorized_chunk_size
num_threshold = self.config.string_to_numeric_threshold
flag_max_distinct = self.config.flag_max_distinct
flag_known = set(
v.lower() for v in self.config.flag_known_values
)
numeric_inferred: List[str] = []
flag_binary: List[str] = []
date_inferred: List[str] = []
pure_categorical: List[str] = []
# Columns pending flag/date analysis after numeric pass
pending_flag_check: List[Tuple[str, int]] = [] # (col, approx_d)
n_chunks = math.ceil(len(non_vec_cols) / chunk_size)
# ── Sub-phase 3A: Numeric Detection ──────────────────────────
for ci in range(n_chunks):
chunk = non_vec_cols[
ci * chunk_size: (ci + 1) * chunk_size
]
print(
f" │ Chunk {ci + 1}/{n_chunks} (3A-Numérico): "
f"{len(chunk)} colunas..."
)
col_keys: List[Tuple[str, str]] = []
try:
agg_exprs = []
for c in chunk:
key = (
c.replace(".", "__")
.replace(" ", "_")
.replace("-", "_")
)
col_keys.append((c, key))
cr = F.col(f"`{c}`")
trimmed = F.trim(cr)
# (1) Non-null, non-empty count
agg_exprs.append(
F.sum(F.when(
cr.isNotNull() & (F.length(trimmed) > 0),
1,
).otherwise(0)).alias(f"{key}___nn")
)
# (2) Castable to double (whole value)
agg_exprs.append(
F.sum(F.when(
cr.isNotNull()
& (F.length(trimmed) > 0)
& trimmed.cast("double").isNotNull(),
1,
).otherwise(0)).alias(f"{key}___cd")
)
# (3) Approximate distinct count
agg_exprs.append(
F.approx_count_distinct(
F.when(
cr.isNotNull() & (F.length(trimmed) > 0),
F.lower(trimmed),
),
).alias(f"{key}___ad")
)
stats = df.agg(*agg_exprs).collect()[0]
for c, key in col_keys:
nn = stats[f"{key}___nn"] or 0
cd = stats[f"{key}___cd"] or 0
ad = stats[f"{key}___ad"] or 0
if nn == 0:
pure_categorical.append(c)
continue
cast_ratio = cd / nn
if cast_ratio >= num_threshold:
numeric_inferred.append(c)
print(
f" │ ✔ {c}: NUMÉRICO INFERIDO "
f"(cast={cast_ratio:.1%}, "
f"distinct≈{ad})"
)
elif ad <= flag_max_distinct:
# Low cardinality → candidate for flag
pending_flag_check.append((c, ad))
else:
# High cardinality, not numeric → categorical
pure_categorical.append(c)
except Exception as e:
print(f" │ ✗ Chunk {ci + 1} (3A) falhou: {e}")
for c, _ in col_keys:
pure_categorical.append(c)
# ── Sub-phase 3B: Flag / Boolean Detection ───────────────────
if pending_flag_check:
print(
f" │\n │ Sub-fase 3B: Verificação de "
f"{len(pending_flag_check)} Flag/Boolean candidatas"
)
# Process in chunks too
flag_chunks = math.ceil(
len(pending_flag_check) / chunk_size,
)
for fci in range(flag_chunks):
fc = pending_flag_check[
fci * chunk_size: (fci + 1) * chunk_size
]
try:
for c, ad in fc:
cr = F.col(f"`{c}`")
trimmed = F.lower(F.trim(cr))
# Collect distinct non-null, non-empty values
distinct_rows = (
df.select(trimmed.alias("v"))
.filter(
F.col("v").isNotNull()
& (F.length(F.col("v")) > 0)
)
.distinct()
.limit(flag_max_distinct + 1)
.collect()
)
distinct_vals = {
row["v"] for row in distinct_rows
}
if (
len(distinct_vals) <= flag_max_distinct
and distinct_vals
and distinct_vals.issubset(flag_known)
):
flag_binary.append(c)
vals_str = ", ".join(
sorted(distinct_vals),
)
print(
f" │ ✔ {c}: FLAG/BOOLEAN "
f"({vals_str})"
)
else:
# Low cardinality but not flag pattern
pure_categorical.append(c)
except Exception as e:
print(
f" │ ✗ Flag chunk {fci + 1} falhou: {e}",
)
for c, _ in fc:
if (
c not in pure_categorical
and c not in flag_binary
):
pure_categorical.append(c)
# ── Sub-phase 3C: Date Detection (on remaining categoricals)
# Only attempt for cols with moderate distinct count
if self.config.skip_date_inference:
print(
f" │\n │ Sub-fase 3C: IGNORADA "
f"(skip_date_inference=True)"
)
else:
date_candidates = [
c for c in pure_categorical
if c not in numeric_inferred
and c not in flag_binary
]
if date_candidates and self.config.date_sample_formats:
print(
f" │\n │ Sub-fase 3C: Teste de data em "
f"{len(date_candidates)} candidatas"
)
tested_dates: List[str] = []
for c in date_candidates:
cr = F.col(f"`{c}`")
trimmed = F.trim(cr)
non_null_filter = (
cr.isNotNull() & (F.length(trimmed) > 0)
)
try:
# Quick row count
nn_row = df.agg(
F.sum(
F.when(non_null_filter, 1).otherwise(0),
).alias("nn"),
).collect()[0]
nn = nn_row["nn"] or 0
if nn == 0:
continue
# Test each format
best_fmt = None
best_ratio = 0.0
for fmt in self.config.date_sample_formats:
parsed = df.agg(
F.sum(F.when(
non_null_filter
& F.to_date(
trimmed, fmt,
).isNotNull(),
1,
).otherwise(0)).alias("dp"),
).collect()[0]
dp = parsed["dp"] or 0
ratio = dp / nn
if ratio > best_ratio:
best_ratio = ratio
best_fmt = fmt
if best_ratio >= 0.95 and best_fmt is not None:
date_inferred.append(c)
tested_dates.append(c)
print(
f" │ ✔ {c}: DATA INFERIDA "
f"(formato='{best_fmt}', "
f"parse={best_ratio:.1%})"
)
except Exception as e:
print(f" │ ✗ Date test '{c}' falhou: {e}")
# Remove date-inferred from pure_categorical
if tested_dates:
pure_categorical = [
c for c in pure_categorical
if c not in tested_dates
]
return numeric_inferred, flag_binary, date_inferred, pure_categorical
def _collect_sample_values(
self,
df: DataFrame,
all_classified: Dict[str, str],
) -> Dict[str, List[str]]:
"""Collects sample non-null values for visual validation.
Uses chunked processing to avoid OOM. For each column, collects
up to ``sample_values_count`` distinct non-null values.
Args:
df (DataFrame): Input DataFrame.
all_classified (Dict[str, str]): Column→type mapping.
Expected Input Schema:
Any DataFrame.
Output Schema Transformation:
Returns dict of column→list of sample value strings.
Memory Implications:
Each chunk collects N values per column via ``limit()``.
N defaults to 5. Safe for any table width.
Exception Handling Protocols:
Per-column failures are silently skipped.
Returns:
Dict[str, List[str]]: Sample values per column.
"""
n_samples = self.config.sample_values_count
cols = list(all_classified.keys())
samples: Dict[str, List[str]] = {}
if not cols:
return samples
# Use a tiny sample of the full DataFrame to avoid full scans
# 0.5% sample with limit provides enough diversity for examples
micro = (
df.select(
[F.col(f"`{c}`").cast("string").alias(c) for c in cols]
)
.sample(withReplacement=False, fraction=0.005, seed=42)
.limit(500)
)
micro_rows = micro.collect()
for c in cols:
try:
vals = set()
for row in micro_rows:
v = row[c]
if v is not None and str(v).strip():
vals.add(str(v))
if len(vals) >= n_samples:
break
samples[c] = list(vals)[:n_samples]
except Exception:
samples[c] = ["<erro>"]
print(f" │ Exemplos coletados para {len(samples)} colunas")
return samples
def _print_classification_report(
self, clf: ColumnClassification, total_cols: int,
) -> None:
"""Prints the final classification summary to stdout.
Args:
clf (ColumnClassification): Classification result.
total_cols (int): Total column count in source DataFrame.
Expected Input Schema:
A populated ``ColumnClassification``.
Output Schema Transformation:
None. Prints to stdout.
Memory Implications:
Negligible.
Exception Handling Protocols:
None.
"""
total_vec = (
len(clf.numeric_vectorized) + len(clf.categorical_vectorized)
)
total_new = sum(clf.vector_lengths.values())
print(f"\n {'=' * 72}")
print(f" {'RELATÓRIO FINAL DE CLASSIFICAÇÃO DE COLUNAS':^72}")
print(f" {'=' * 72}")
print(f" │ Total de Colunas Analisadas: {total_cols:>6}")
print(f" │ {'─' * 50}")
print(f" │ Numéricas Nativas (schema): {len(clf.numeric):>6}")
print(f" │ Numéricas Inferidas (string→dbl): {len(clf.numeric_inferred):>6}")
print(f" │ Flag / Boolean: {len(clf.flag_binary):>6}")
print(f" │ Categóricas (string puro): {len(clf.categorical):>6}")
print(f" │ Numéricas Vetorizadas (#): {len(clf.numeric_vectorized):>6}")
print(f" │ Categóricas Vetorizadas (#): {len(clf.categorical_vectorized):>6}")
print(f" │ Temporais (date/timestamp): {len(clf.temporal):>6}")
print(f" │ Datas Inferidas (string→date): {len(clf.date_inferred):>6}")
print(f" │ Excluídas (complex/binary): {len(clf.excluded):>6}")
print(f" │ {'─' * 50}")
print(f" │ Total Vetorizadas: {total_vec:>6}")
print(f" │ Sub-colunas a gerar (explosão): {total_new:>6}")
print(f" {'=' * 72}")
def _print_detailed_sample_report(
self, clf: ColumnClassification,
) -> None:
"""Prints a detailed per-type, per-column sample value report.
Designed for the user to visually validate that each column
was correctly classified by inspecting real data samples.
Args:
clf (ColumnClassification): Classification result.
Expected Input Schema:
``clf.sample_values`` must be populated.
Output Schema Transformation:
None. Prints to stdout.
Memory Implications:
Negligible.
Exception Handling Protocols:
None.
"""
sv = clf.sample_values
if not sv:
return
type_groups = [
("NUMÉRICAS NATIVAS", clf.numeric),
("NUMÉRICAS INFERIDAS (string→double)", clf.numeric_inferred),
("FLAG / BOOLEAN", clf.flag_binary),
("CATEGÓRICAS", clf.categorical),
("NUMÉRICAS VETORIZADAS (#)", clf.numeric_vectorized),
("CATEGÓRICAS VETORIZADAS (#)", clf.categorical_vectorized),
("TEMPORAIS", clf.temporal),
("DATAS INFERIDAS (string→date)", clf.date_inferred),
("EXCLUÍDAS", clf.excluded),
]
print(f"\n {'=' * 72}")
print(f" {'EXEMPLOS DE VALORES POR CATEGORIA (para validação visual)':^72}")
print(f" {'=' * 72}")
for type_label, col_list in type_groups:
if not col_list:
continue
print(f"\n ┌── {type_label} ({len(col_list)} colunas)")
# Show up to 20 columns per type to avoid flooding
display_cols = col_list[:20]
for c in display_cols:
vals = sv.get(c, ["<sem dados>"])
vals_str = " | ".join(str(v) for v in vals)
# Truncate very long displays
if len(vals_str) > 90:
vals_str = vals_str[:87] + "..."
print(f" │ {c}")
print(f" │ → [{vals_str}]")
if len(col_list) > 20:
print(
f" │ ... e mais {len(col_list) - 20} "
f"colunas (ver Delta para lista completa)"
)
print(f" └{'─' * 71}")
def _persist_classification(
self, clf: ColumnClassification,
) -> None:
"""Persists the classification report as a Delta table.
Args:
clf (ColumnClassification): Classification result.
Expected Input Schema:
A populated ``ColumnClassification``.
Output Schema Transformation:
Creates/overwrites Delta table:
``{dest_schema}.tb_column_classification_{cart}_{YYYYMM}``.
Includes ``sample_values`` serialised as pipe-separated string.
Memory Implications:
Creates a small DataFrame on the Driver (~500 rows max).
Exception Handling Protocols:
Catches write failures and logs a warning.
"""
rows: List[Dict[str, Any]] = []
ts = datetime.now().isoformat()
cart = self.config.carteira_ifrs
type_map = [
(clf.numeric, "numeric"),
(clf.numeric_inferred, "numeric_inferred"),
(clf.flag_binary, "flag_binary"),
(clf.categorical, "categorical"),
(clf.numeric_vectorized, "numeric_vectorized"),
(clf.categorical_vectorized, "categorical_vectorized"),
(clf.temporal, "temporal"),
(clf.date_inferred, "date_inferred"),
(clf.excluded, "excluded"),
]
for col_list, detected_type in type_map:
for c in col_list:
sample_vals = clf.sample_values.get(c, [])
rows.append({
"column_name": c,
"detected_type": detected_type,
"vector_length": clf.vector_lengths.get(c),
"sample_values": " | ".join(
str(v) for v in sample_vals
),
"carteira": cart,
"analyzed_at": ts,
})
if not rows:
return
try:
cart_safe = cart.lower().replace(" ", "_")
date_suffix = datetime.now().strftime("%Y%m")
table_name = (
f"tb_column_classification_{cart_safe}_{date_suffix}"
)
full_table = f"{self.config.dest_schema}.{table_name}"
report_df = self.spark.createDataFrame(rows)
report_df.write.format("delta").mode("overwrite").option(
"overwriteSchema", "true",
).saveAsTable(full_table)
print(
f"\n ✔ Classificação salva: {full_table} "
f"→ {len(rows)} registros"
)
except Exception as e:
print(f" ⚠ Falha ao persistir classificação: {e}")
# ╔═════════════════════════════════════════════════════════════════════════╗
# ║ 2D. VECTORIZED COLUMN HANDLER — HASH-SEPARATED EXPLOSION ║
# ╚═════════════════════════════════════════════════════════════════════════╝
class VectorizedColumnHandler:
"""Distributed engine for exploding ``#``-separated vectorised columns.
Takes the ``ColumnClassification`` output from ``ColumnTypeAnalyzer``
and creates individual typed sub-columns for each detected vectorised
column.
Example::
Column ``pd_term`` with value ``"0.01#0.02#0.03"``
(max_length=3, type=numeric) produces:
pd_term_0 = 0.01 (double)
pd_term_1 = 0.02 (double)
pd_term_2 = 0.03 (double)
OOM Prevention:
- Processes in chunks of ``vectorized_explode_chunk_size``
(default: 5) vectorised columns per iteration.
- Uses ``localCheckpoint(eager=True)`` between chunks to
truncate the Spark DAG lineage and prevent
StackOverflowError on deeply nested plans.
- Each chunk adds new columns via a single
``df.select("*", ...)`` call to minimise Catalyst plan depth.
Attributes:
spark (SparkSession): Active session.
config (SamplerConfig): Suite configuration.
"""
def __init__(
self, spark: SparkSession, config: SamplerConfig,
) -> None:
"""Initializes the handler.
Args:
spark (SparkSession): Active Spark session.
config (SamplerConfig): Suite configuration.
Expected Input Schema:
None. Configuration only.
Output Schema Transformation:
None.
Memory Implications:
Negligible.
Exception Handling Protocols:
None.
"""
self.spark = spark
self.config = config
def explode_all(
self, df: DataFrame, clf: ColumnClassification,
) -> DataFrame:
"""Explodes all vectorised columns into individual sub-columns.
Args:
df (DataFrame): Input DataFrame with original vectorised cols.
clf (ColumnClassification): Output from ColumnTypeAnalyzer.
Expected Input Schema:
DataFrame containing all vectorised columns listed in ``clf``.
Output Schema Transformation:
Adds ``{col}_{i}`` for ``i ∈ [0, max_len)`` per vectorised col.
Original columns are PRESERVED (not dropped).
Numeric vectorised sub-columns are cast to ``double``.
Categorical vectorised sub-columns are trimmed strings
(empty strings → ``null``).
Memory Implications:
Creates ``sum(vector_lengths)`` additional columns.
Processed in chunks of ``explode_chunk_size`` with
``localCheckpoint`` between chunks to truncate lineage.
Exception Handling Protocols:
Per-chunk failures are caught and logged. Successful chunks
are retained; failed columns are skipped.
Returns:
DataFrame: Input DataFrame augmented with sub-columns.
"""
all_vec = (
clf.numeric_vectorized + clf.categorical_vectorized
)
if not all_vec:
print(" ├── Nenhuma coluna vetorizada para explodir.")
return df
sep = self.config.vectorized_separator
escaped_sep = re.escape(sep)
max_cap = self.config.vectorized_max_elements
budget = self.config.vectorized_max_subcols_per_chunk
# ── Apply optional element cap & compute effective lengths ──
effective_lengths: Dict[str, int] = {}
for c in all_vec:
detected = clf.vector_lengths.get(c, 1)
if max_cap is not None and detected > max_cap:
effective_lengths[c] = max_cap
print(
f" ⚠ {c}: vetor com {detected} elementos "
f"→ limitado a {max_cap} (vectorized_max_elements)"
)
else:
effective_lengths[c] = detected
total_new = sum(effective_lengths.values())
# ── Build adaptive chunks (budget = max sub-cols per chunk) ──
chunks: List[List[str]] = []
current_chunk: List[str] = []
current_budget = 0
for c in all_vec:
col_len = effective_lengths[c]
# If single column already exceeds budget, isolate it
if col_len >= budget:
if current_chunk:
chunks.append(current_chunk)
chunks.append([c])
current_chunk = []
current_budget = 0
elif current_budget + col_len > budget:
# Flush current chunk, start new one
chunks.append(current_chunk)
current_chunk = [c]
current_budget = col_len
else:
current_chunk.append(c)
current_budget += col_len
if current_chunk:
chunks.append(current_chunk)
n_chunks = len(chunks)
print(f"\n ┌── EXPLOSÃO DE COLUNAS VETORIZADAS")
print(f" │ Colunas vetorizadas: {len(all_vec)}")
print(f" │ Sub-colunas a gerar: {total_new}")
print(
f" │ Chunks adaptativos: {n_chunks} "
f"(budget ≤ {budget} sub-cols/chunk)"
)
if max_cap is not None:
print(f" │ Cap por coluna: {max_cap} elementos")
for ci, chunk in enumerate(chunks):
chunk_subcols = sum(effective_lengths[c] for c in chunk)
print(f" │")
print(
f" ├── Chunk {ci + 1}/{n_chunks}: "
f"{len(chunk)} col(s), {chunk_subcols} sub-colunas..."
)
try:
new_col_exprs = []
for col_name in chunk:
eff_len = effective_lengths[col_name]
is_numeric = (
col_name in clf.numeric_vectorized
)
split_expr = F.split(
F.col(f"`{col_name}`"), escaped_sep,
)
for j in range(eff_len):
sub_name = f"{col_name}_{j}"
# element_at is 1-indexed; returns null OOB
elem = F.element_at(split_expr, j + 1)
if is_numeric:
expr = elem.cast("double")
else:
# Trim whitespace; empty → null
trimmed = F.trim(elem)
expr = F.when(
F.length(trimmed) > 0, trimmed,
)
new_col_exprs.append(expr.alias(sub_name))
label = "double" if is_numeric else "string"
print(
f" │ ✔ {col_name} → {eff_len} ({label})"
)
# Single select to add all new columns
df = df.select("*", *new_col_exprs)
# Truncate lineage between chunks
if ci < n_chunks - 1:
df = df.localCheckpoint(eager=True)
print(
f" │ ↻ Lineage truncado "
f"(localCheckpoint)"
)
except Exception as e:
print(f" │ ✗ Chunk {ci + 1} falhou: {e}")
print(
f" └── ✔ Explosão completa. "
f"Schema: {len(df.columns)} colunas totais"
)
return df
# ╔═════════════════════════════════════════════════════════════════════════╗
# ║ 3. CASTING & FILTERING ║
# ╚═════════════════════════════════════════════════════════════════════════╝
def cast_and_filter(
spark: SparkSession,
source_table: str,
carteira_ifrs: str,
reporting_date_start: Optional[str] = None,
castings: Dict[str, str] = DICT_CASTINGS,
) -> DataFrame:
"""
Reads the source table, applies column casts, filters by carteira,
and optionally applies a temporal lower-bound filter on reporting_date.
Args:
spark (SparkSession): Active Spark session.
source_table (str): Fully-qualified table name.
carteira_ifrs (str): Value to filter ``carteira_ifrs`` column.
reporting_date_start (Optional[str]): ISO date string
(e.g., ``"2022-01-01"``). If provided, only rows with
``reporting_date >= reporting_date_start`` are kept.
Enables predicate pushdown on the date partition.
castings (Dict[str, str]): Column-name → Spark-type mapping.
Expected Input Schema:
Any DataFrame readable via ``spark.table()``.
Output Schema Transformation:
Returns a DataFrame filtered to the given ``carteira_ifrs`` value
(and optionally ``reporting_date >= start``) with all columns in
``castings`` cast to their target types. Columns not present in
the source are silently skipped.
Memory Implications:
Predicate pushdown on ``carteira_ifrs`` + ``reporting_date``
reduces the partition scan substantially (typically 5-30% of
the full multi-year dataset).
Exception Handling Protocols:
Logs a warning for columns listed in ``castings`` but absent from
the source schema and continues without them.
Returns:
DataFrame: Filtered and cast DataFrame.
"""
df = spark.table(source_table)
existing_cols = set(df.columns)
for col_name, col_type in castings.items():
if col_name in existing_cols:
df = df.withColumn(col_name, F.col(col_name).cast(col_type))
else:
print(f" ⚠ Coluna '{col_name}' ausente na origem — ignorada.")
# ── Carteira filter ──
df = df.filter(F.col("carteira_ifrs") == carteira_ifrs)
# ── Temporal lower-bound filter ──
if reporting_date_start is not None:
df = df.filter(F.col("reporting_date") >= F.lit(reporting_date_start).cast("date"))
print(f" ✔ Filtro temporal: reporting_date >= '{reporting_date_start}'")
return df
# ╔═════════════════════════════════════════════════════════════════════════╗
# ║ 4. GENERIC NEYMAN VOLATILITY SAMPLER (REUSABLE CORE) ║
# ╚═════════════════════════════════════════════════════════════════════════╝
def _compute_neyman_sample(
df: DataFrame,
group_cols: List[str],
target_col: str,
total_budget: int,
min_per_stratum: int,
use_ntile: bool = False,
ntile_col: Optional[str] = None,
n_tiles: int = 10,
) -> DataFrame:
"""
Core engine: builds strata from ``group_cols``, computes variance of
``target_col`` per stratum, allocates via RNABOX Neyman, and draws.
This function is intentionally generic so it can power multiple sample
strategies (A1, A2, A3) by varying ``group_cols`` and ``use_ntile``.
Args:
df (DataFrame): Already filtered/cast population DataFrame.
group_cols (List[str]): Columns whose categorical cross-product
defines the strata (e.g., ``["stage"]``).
target_col (str): Column whose variance drives Neyman weights.
total_budget (int): Total sample size n.
min_per_stratum (int): RNABOX floor L_h.
use_ntile (bool): If True, adds an ``ntile`` column within each
combination of ``group_cols`` ordered by ``ntile_col``.
ntile_col (Optional[str]): Column to order for ntile computation.
n_tiles (int): Number of quantile buckets (default 10 = deciles).
Expected Input Schema:
DataFrame must contain all columns in ``group_cols``, the
``target_col``, and ``ntile_col`` (when ``use_ntile=True``).
Output Schema Transformation:
Returns a sampled DataFrame with all original columns plus:
- ``stratum_id`` (StringType): Composite stratum key.
- ``_vol_decile`` (IntegerType): Only when ``use_ntile=True``.
Memory Implications:
- One ``groupBy`` aggregation (bounded by stratum cardinality).
- RNABOX runs on the driver over ~30-100 rows. Negligible.
- ``sampleBy`` is a single distributed scan with per-partition
Bernoulli sampling.
Exception Handling Protocols:
Falls back to proportional allocation (floor=1) if the budget
cannot satisfy all floor constraints.
Returns:
DataFrame: Neyman-optimal stratified sample.
"""
# ── Step 1: Optionally add decile column ──
if use_ntile and ntile_col is not None:
w_ntile = Window.partitionBy(*group_cols).orderBy(ntile_col)
df = df.withColumn("_vol_decile", F.ntile(n_tiles).over(w_ntile))
strata_key_cols = group_cols + ["_vol_decile"]
else:
strata_key_cols = list(group_cols)
# ── Step 2: Build stratum_id ──
concat_parts = []
for c in strata_key_cols:
concat_parts.append(F.coalesce(F.col(c).cast("string"), F.lit("null")))
concat_parts.append(F.lit("_"))
# Remove trailing separator
concat_parts = concat_parts[:-1]
df = df.withColumn("stratum_id", F.concat(*concat_parts))
# ── Step 3: Per-stratum statistics ──
stats_df = df.groupBy("stratum_id").agg(
F.count("*").alias("N_h"),
F.stddev(target_col).alias("S_h"),
)
stats_rows = stats_df.collect()
strata_stats: Dict[str, Tuple[int, float]] = {}
for row in stats_rows:
sid = row["stratum_id"]
nh = row["N_h"]
sh = row["S_h"] if row["S_h"] is not None and row["S_h"] > 0 else 1e-9
strata_stats[sid] = (nh, sh)
if not strata_stats:
print(" ⚠ Nenhum estrato encontrado — retornando DataFrame vazio.")
return df.limit(0)
# Adjust budget if it exceeds population
total_pop = sum(nh for nh, _ in strata_stats.values())
effective_budget = min(total_budget, total_pop)
# ── Step 4: RNABOX Allocation ──
n_strata = len(strata_stats)
min_required = n_strata * min_per_stratum
if effective_budget < min_required:
# Fallback: floor = 1
allocator = RNABOXAllocator(min_per_stratum=1)
else:
allocator = RNABOXAllocator(min_per_stratum=min_per_stratum)
allocations = allocator.allocate(strata_stats, effective_budget)
# ── Step 5: Convert to fractions and draw ──
fractions: Dict[str, float] = {}
for sid, n_alloc in allocations.items():
nh_pop = strata_stats[sid][0]
fractions[sid] = min(n_alloc / nh_pop, 1.0)
sampled = df.stat.sampleBy("stratum_id", fractions, seed=42)
# Log allocation summary
print(f" ├── Estratos: {n_strata} | Pop: {total_pop:,} | Budget: {effective_budget:,}")
for sid in sorted(allocations.keys()):
nh, sh = strata_stats[sid]
n_alloc = allocations[sid]
print(f" │ {sid:30s} N_h={nh:>8,} S_h={sh:>12.2f} n_h={n_alloc:>6,}")
return sampled
# ╔═════════════════════════════════════════════════════════════════════════╗
# ║ 5. PSI CALCULATOR (100% PYSPARK) ║
# ╚═════════════════════════════════════════════════════════════════════════╝
def compute_psi_spark(
reference_df: DataFrame,
sample_df: DataFrame,
col_name: str,
n_bins: int = 10,
) -> Tuple[float, str]:
"""
Computes the Population Stability Index (PSI) between two DataFrames
for a given continuous column, 100% distributed in PySpark.
Algorithm:
1. Compute decile boundaries from the reference DataFrame using
``approxQuantile``.
2. Bucket both DataFrames into ``n_bins`` bins.
3. Compute bin proportions for each.
4. PSI = Σ (p_i - q_i) × ln(p_i / q_i)
Args:
reference_df (DataFrame): Population (source) DataFrame.
sample_df (DataFrame): Sample DataFrame to validate.
col_name (str): Column to compute PSI on.
n_bins (int): Number of bins for discretisation.
Expected Input Schema:
Both DataFrames must contain ``col_name`` as a numeric column.
Output Schema Transformation:
Returns a tuple ``(psi_value, signal)`` where ``signal`` is
one of ``"🟢 VERDE"``, ``"🟡 AMARELO"``, ``"🔴 VERMELHO"``.
Memory Implications:
``approxQuantile`` is distributed (T-Digest). The bucketing
and proportion computation use two groupBy aggregations with
``n_bins`` groups each — negligible overhead.
Exception Handling Protocols:
Returns ``(0.0, "🟢 VERDE")`` if either DataFrame is empty or
the column has zero variance.
Returns:
Tuple[float, str]: (PSI value, traffic-light signal).
"""
# Guard: empty inputs
ref_count = reference_df.select(col_name).filter(F.col(col_name).isNotNull()).count()
spl_count = sample_df.select(col_name).filter(F.col(col_name).isNotNull()).count()
if ref_count == 0 or spl_count == 0:
return 0.0, "🟢 VERDE"
# Step 1: Compute quantile boundaries from reference
quantiles = [i / n_bins for i in range(1, n_bins)]
boundaries = reference_df.approxQuantile(col_name, quantiles, 0.01)
if not boundaries or len(set(boundaries)) < 2:
return 0.0, "🟢 VERDE"
# Ensure unique sorted boundaries
boundaries = sorted(set(boundaries))
# Step 2: Bucket function
def _assign_bucket(df: DataFrame, col: str, bounds: List[float]) -> DataFrame:
"""Assigns each row to a bin based on quantile boundaries.
Args:
df (DataFrame): Input DataFrame.
col (str): Column to bucket.
bounds (List[float]): Sorted boundary values.
Expected Input Schema:
DataFrame with numeric ``col``.
Output Schema Transformation:
Adds ``_psi_bucket`` (IntegerType) column.
Memory Implications:
Single pass with chained ``F.when`` — no shuffle.
Exception Handling Protocols:
Values below min boundary → bucket 0.
Values above max boundary → last bucket.
"""
bucket_expr = F.lit(0)
for i, b in enumerate(bounds):
bucket_expr = F.when(
F.col(col) >= F.lit(b), F.lit(i + 1)
).otherwise(bucket_expr)
return df.withColumn("_psi_bucket", bucket_expr)
ref_bucketed = _assign_bucket(reference_df, col_name, boundaries)
spl_bucketed = _assign_bucket(sample_df, col_name, boundaries)
# Step 3: Compute proportions
ref_props = (
ref_bucketed
.groupBy("_psi_bucket")
.agg(F.count("*").alias("ref_count"))
.withColumn("ref_prop", F.col("ref_count") / F.lit(ref_count))
)
spl_props = (
spl_bucketed
.groupBy("_psi_bucket")
.agg(F.count("*").alias("spl_count"))
.withColumn("spl_prop", F.col("spl_count") / F.lit(spl_count))
)
# Step 4: Join and compute PSI components
joined = ref_props.join(spl_props, on="_psi_bucket", how="outer")
joined = joined.fillna({"ref_prop": 1e-10, "spl_prop": 1e-10})
# Floor small proportions to avoid log(0)
joined = joined.withColumn(
"ref_prop", F.greatest(F.col("ref_prop"), F.lit(1e-10))
).withColumn(
"spl_prop", F.greatest(F.col("spl_prop"), F.lit(1e-10))
)
# PSI per bin = (p_i - q_i) * ln(p_i / q_i)
joined = joined.withColumn(
"_psi_component",
(F.col("spl_prop") - F.col("ref_prop"))
* F.log(F.col("spl_prop") / F.col("ref_prop")),
)
psi_row = joined.agg(F.sum("_psi_component").alias("psi")).collect()
psi_value = psi_row[0]["psi"] if psi_row and psi_row[0]["psi"] is not None else 0.0
psi_value = abs(psi_value)
# Traffic light
if psi_value < 0.1:
signal = "🟢 VERDE"
elif psi_value <= 0.2:
signal = "🟡 AMARELO"
else:
signal = "🔴 VERMELHO"
return psi_value, signal
# ╔═════════════════════════════════════════════════════════════════════════╗
# ║ 6. VALIDATION ORCHESTRATOR ║
# ╚═════════════════════════════════════════════════════════════════════════╝
def validate_sample(
reference_df: DataFrame,
sample_df: DataFrame,
sample_name: str,
config: SamplerConfig,
) -> DataFrame:
"""
Runs PSI and PRS validation on a generated sample against the source
population and prints a semaphore report.
Args:
reference_df (DataFrame): Full population DataFrame.
sample_df (DataFrame): The generated sample to validate.
sample_name (str): Human-readable name for logging.
config (SamplerConfig): Suite configuration.
Expected Input Schema:
Both DataFrames must contain ``config.target_col`` and all
columns listed in ``config.validation_cols``.
Output Schema Transformation:
Returns a summary DataFrame with columns:
- ``sample_name`` (StringType)
- ``column`` (StringType)
- ``psi_value`` (DoubleType)
- ``psi_signal`` (StringType)
- ``prs_value`` (DoubleType)
- ``prs_stable`` (StringType)
Memory Implications:
Runs ``approxQuantile`` (T-Digest) per column. With ~3 columns
and 10 bins, this is ~30 quantile computations. Negligible.
Exception Handling Protocols:
Columns missing from either DataFrame are skipped with a warning.
Returns:
DataFrame: Validation results as a Spark DataFrame.
"""
spark = config.spark
cols_to_validate = [config.target_col] + config.validation_cols
results: List[Dict[str, Any]] = []
prs_validator = PRSValidator(
epsilon=config.prs_epsilon,
alpha=config.prs_alpha,
n_bins=config.psi_n_bins,
)
print(f"\n ┌── Validação: {sample_name}")
print(f" │ Pop: {reference_df.count():,} | Sample: {sample_df.count():,}")
for col_name in cols_to_validate:
if col_name not in reference_df.columns or col_name not in sample_df.columns:
print(f" │ ⚠ Coluna '{col_name}' ausente — ignorada.")
continue
# PSI (100% PySpark)
psi_val, psi_signal = compute_psi_spark(
reference_df, sample_df, col_name, config.psi_n_bins,
)
# PRS (uses proportions computed in PySpark, Chi-Square in NumPy)
quantiles = [i / config.psi_n_bins for i in range(1, config.psi_n_bins)]
boundaries = reference_df.approxQuantile(col_name, quantiles, 0.01)
boundaries = sorted(set(boundaries)) if boundaries else []
prs_val = 0.0
prs_stable = "N/A"
if len(boundaries) >= 2:
import numpy as np
def _get_proportions(df: DataFrame, col: str, bounds: List[float]) -> "np.ndarray":
"""Computes bin proportions for PRS using PySpark bucketing.
Args:
df (DataFrame): Input DataFrame.
col (str): Column to bucket.
bounds (List[float]): Quantile boundaries.
Expected Input Schema:
DataFrame with numeric ``col``.
Output Schema Transformation:
Returns np.ndarray of proportions (shape n_bins+1).
Memory Implications:
One groupBy with n_bins groups. Negligible.
Exception Handling Protocols:
Returns uniform array if bucketing fails.
"""
bucket_expr = F.lit(0)
for i, b in enumerate(bounds):
bucket_expr = F.when(
F.col(col) >= F.lit(b), F.lit(i + 1)
).otherwise(bucket_expr)
bucketed = df.withColumn("_b", bucket_expr)
total = bucketed.count()
if total == 0:
n = len(bounds) + 1
return np.ones(n) / n
counts = (
bucketed.groupBy("_b")
.agg(F.count("*").alias("cnt"))
.collect()
)
n = len(bounds) + 1
arr = np.zeros(n)
for row in counts:
idx = row["_b"] if row["_b"] is not None else 0
if 0 <= idx < n:
arr[idx] = row["cnt"]
return arr / arr.sum() if arr.sum() > 0 else np.ones(n) / n
ref_props = _get_proportions(reference_df, col_name, boundaries)
spl_props = _get_proportions(sample_df, col_name, boundaries)
spl_n = sample_df.filter(F.col(col_name).isNotNull()).count()
if spl_n > 0:
prs_result = prs_validator.evaluate(ref_props, spl_props, spl_n)
prs_val = prs_result["prs_value"]
prs_stable = "STABLE" if prs_result["is_stable"] else "DRIFT"
print(
f" │ {col_name:35s} PSI={psi_val:.4f} {psi_signal}"
f" | PRS={prs_val:.4f} [{prs_stable}]"
)
results.append({
"sample_name": sample_name,
"column": col_name,
"psi_value": round(psi_val, 6),
"psi_signal": psi_signal,
"prs_value": round(prs_val, 6),
"prs_stable": prs_stable,
})
print(f" └───────────────────────────────────────────────────")
if spark is not None and results:
return spark.createDataFrame(results)
return None
# ╔═════════════════════════════════════════════════════════════════════════╗
# ║ 7. IFRS 9 SAMPLER SUITE (ORCHESTRATOR CLASS) ║
# ╚═════════════════════════════════════════════════════════════════════════╝
class IFRS9SamplerSuite:
"""
End-to-end orchestrator that generates all static samples, validates
them, and persists to Delta in ``prd.sand_crc_estudos_ifrs9``.
Workflow (modular — each step can be called independently):
1. ``load_source()`` — Load and prepare the source data.
2. ``run_data_quality_profiling()`` — Profile all columns.
3. ``compute_minimum_sample_size()`` — Probabilistic floor.
4. ``generate_all_samples()`` — Produce A1/A2/A3/B/B1/B2/C/D/E.
5. ``validate_all()`` — Run PSI + PRS on every sample.
6. ``persist_all()`` — Write each to Delta with overwrite.
Attributes:
config (SamplerConfig): Suite configuration.
source_df (Optional[DataFrame]): Cached source population.
profiling_df (Optional[DataFrame]): Data Quality profiling.
profiler (Optional[DataQualityProfiler]): Profiler instance.
samples (Dict[str, DataFrame]): Generated samples by name.
validation_reports (List[DataFrame]): Per-sample validation.
"""
def __init__(self, config: SamplerConfig) -> None:
"""
Initializes the suite.
Args:
config (SamplerConfig): Fully populated configuration.
Expected Input Schema:
None. Configuration only.
Output Schema Transformation:
None.
Memory Implications:
Negligible. Stores references and scalar parameters.
Exception Handling Protocols:
Raises ValueError if ``config.spark`` is None.
"""
if config.spark is None:
raise ValueError("SamplerConfig.spark must be set.")
self.config = config
self.source_df: Optional[DataFrame] = None
self.profiling_df: Optional[DataFrame] = None
self.profiler: Optional[DataQualityProfiler] = None
self.samples: Dict[str, DataFrame] = {}
self.validation_reports: List[DataFrame] = []
self.column_classification: Optional[ColumnClassification] = None
# ── 7.1 Load ──────────────────────────────────────────────────────
def load_source(self) -> DataFrame:
"""
Reads, casts, and filters the source table. Caches result.
Applies the ``reporting_date_start`` temporal filter if configured.
Expected Input Schema:
Raw table from Unity Catalog.
Output Schema Transformation:
Cast columns per ``DICT_CASTINGS``, filtered to carteira
and optionally by ``reporting_date >= start``.
Memory Implications:
Caches the filtered DataFrame. Size depends on portfolio
and date range.
Exception Handling Protocols:
Propagates Spark read errors.
Returns:
DataFrame: Filtered and cast source population.
"""
cart = self.config.carteira_ifrs
date_start = self.config.reporting_date_start
print(f"\n{'='*72}")
print(f" IFRS9 Sampler Suite — Carteira: {cart}")
print(f" Origem: {self.config.source_table}")
print(f" Destino: {self.config.dest_schema}")
if date_start:
print(f" Data Início: {date_start}")
print(f"{'='*72}")
if self.config.enable_delta_checkpoints:
cart_slug = cart.lower().replace(" ", "_")
table_name = f"{self.config.checkpoint_prefix}source_{cart_slug}"
full_path = f"{self.config.dest_schema}.{table_name}"
try:
chkpt_df = self.config.spark.table(full_path)
print(f" ✔ Checkpoint source encontrado: {full_path} (Bypass de carga!)")
chkpt_df = chkpt_df.localCheckpoint(eager=False)
chkpt_df.persist(self._get_storage_level())
self.source_df = chkpt_df
n = self.source_df.count()
print(f" ✔ Origem recuperada do checkpoint: {n:,} registros para '{cart}'")
return self.source_df
except Exception:
pass # Checkpoint does not exist, compute it
self.source_df = cast_and_filter(
spark=self.config.spark,
source_table=self.config.source_table,
carteira_ifrs=cart,
reporting_date_start=date_start,
)
# ── Repartition for cluster parallelism ──
n_parts = self.config.n_repartition
if n_parts is None:
try:
sc = self.config.spark.sparkContext
n_parts = sc.defaultParallelism
except Exception:
n_parts = 96 # 16 cores × 6 workers
self.source_df = self.source_df.repartition(n_parts)
# ── Save as Checkpoint ──
if self.config.enable_delta_checkpoints:
self.source_df = self._checkpoint_dataframe(self.source_df, "source")
else:
sl = self._get_storage_level()
self.source_df.persist(sl)
n = self.source_df.count()
print(f" ✔ Origem carregada: {n:,} registros para '{cart}'")
print(
f" Partições: {n_parts} | "
f"Storage: {self.config.storage_level} | "
f"Colunas: {len(self.source_df.columns)}"
)
return self.source_df
# ── 7.1-mem Memory Utilities ───────────────────────────────────
def _get_storage_level(self) -> StorageLevel:
"""Resolves the configured storage level string to a StorageLevel.
Args:
None.
Expected Input Schema:
``self.config.storage_level`` string.
Output Schema Transformation:
Returns ``pyspark.StorageLevel``.
Memory Implications:
None.
Exception Handling Protocols:
Falls back to ``MEMORY_AND_DISK`` on unknown string.
Returns:
StorageLevel: Resolved level.
"""
mapping = {
"MEMORY_ONLY": StorageLevel.MEMORY_ONLY,
"MEMORY_AND_DISK": StorageLevel.MEMORY_AND_DISK,
"DISK_ONLY": StorageLevel.DISK_ONLY,
"MEMORY_AND_DISK_SER": StorageLevel.MEMORY_AND_DISK, # Fallback mapping
}
return mapping.get(
self.config.storage_level.upper(),
StorageLevel.MEMORY_AND_DISK,
)
# ── 7.1-chkpt Checkpointing Utilities ──────────────────────────
def _checkpoint_dataframe(
self, df: DataFrame, step_name: str,
) -> DataFrame:
"""Saves a DataFrame to Delta as a checkpoint, or loads it if it exists.
Args:
df (DataFrame): The DataFrame to checkpoint.
step_name (str): Identifier for this step (e.g., "source", "exploded").
Returns:
DataFrame: The DataFrame loaded from the Delta checkpoint.
"""
if not self.config.enable_delta_checkpoints:
return df
cart = self.config.carteira_ifrs.lower().replace(" ", "_")
table_name = f"{self.config.checkpoint_prefix}{step_name}_{cart}"
full_path = f"{self.config.dest_schema}.{table_name}"
try:
# Try to read the checkpoint
spark = self.config.spark
# If table doesn't exist, this will throw an exception
chkpt_df = spark.table(full_path)
print(f" ✔ Checkpoint encontrado: {full_path} (Restauração rápida!)")
# Force lineage break and persist again
chkpt_df = chkpt_df.localCheckpoint(eager=False)
chkpt_df.persist(self._get_storage_level())
return chkpt_df
except Exception:
# Table doesn't exist, we need to create it
print(f" ├── Escrevendo checkpoint Delta: {full_path} ...")
try:
df.write.format("delta").mode("overwrite").option(
"overwriteSchema", "true",
).saveAsTable(full_path)
# Unpersist the old in-memory dataframe since we wrote it
df.unpersist()
# Load it back to break lineage completely
chkpt_df = self.config.spark.table(full_path)
chkpt_df = chkpt_df.localCheckpoint(eager=False)
chkpt_df.persist(self._get_storage_level())
print(f" ✔ Checkpoint salvo com sucesso.")
return chkpt_df
except Exception as e:
print(f" ✗ Falha ao salvar checkpoint '{full_path}': {e}")
return df
def _recover_classification_from_delta(self) -> Optional['ColumnClassification']:
"""Attempts to recover the classification from its Delta table."""
if not self.config.enable_delta_checkpoints:
return None
cart = self.config.carteira_ifrs.lower().replace(" ", "_")
dt_str = datetime.now().strftime("%Y%m")
table_name = f"tb_column_classification_{cart}_{dt_str}"
full_table = f"{self.config.dest_schema}.{table_name}"
try:
df_class = self.config.spark.table(full_table)
print(f" ✔ Classificação recuperada do checkpoint ({full_table})!")
# Reconstruct the classification object
rows = df_class.collect()
numeric = []
string = []
boolean = []
temporal = []
complex_type = []
num_vec = []
cat_vec = []
pure_cat = []
num_inf = []
flag_bin = []
date_inf = []
vec_len = {}
for r in rows:
col = r["column"]
cat = r["inferred_category"]
if cat == "numeric": numeric.append(col)
elif cat == "string": string.append(col)
elif cat == "boolean": boolean.append(col)
elif cat == "temporal": temporal.append(col)
elif cat == "complex": complex_type.append(col)
elif cat == "numeric_vectorized": num_vec.append(col)
elif cat == "categorical_vectorized": cat_vec.append(col)
elif cat == "categorical": pure_cat.append(col)
elif cat == "numeric_inferred": num_inf.append(col)
elif cat == "flag_binary": flag_bin.append(col)
elif cat == "date_inferred": date_inf.append(col)
if cat in ("numeric_vectorized", "categorical_vectorized") and r["max_vector_length"]:
vec_len[col] = int(r["max_vector_length"])
return ColumnClassification(
numeric=numeric, string=string, boolean=boolean, temporal=temporal, complex_type=complex_type,
numeric_vectorized=num_vec, categorical_vectorized=cat_vec, pure_categorical=pure_cat,
numeric_inferred=num_inf, flag_binary=flag_bin, date_inferred=date_inf,
vector_lengths=vec_len, sample_values={}
)
except Exception:
return None
def release_memory(self, full: bool = False) -> None:
"""Releases cached DataFrames and runs JVM + Python GC.
Call this between heavy cells to reclaim executor memory.
Args:
full (bool): If True, also clears ALL Spark caches
(including source_df). Use only when you know what
you're doing (e.g., between sampling and validation).
Expected Input Schema:
None.
Output Schema Transformation:
None.
Memory Implications:
Reclaims cached memory. After ``full=True``, ``source_df``
will need to be re-read if accessed again.
Exception Handling Protocols:
Silently ignores all errors.
"""
try:
if full:
self.config.spark.catalog.clearCache()
print(" ✔ spark.catalog.clearCache() executado")
# Force Python GC
gc.collect()
# Force JVM GC (best-effort)
self.config.spark.sparkContext._jvm.System.gc()
print(" ✔ GC executado (Python + JVM)")
except Exception:
gc.collect()
print(" ✔ GC executado (Python)")
# ── 7.1a-bis Processing Per-Sample (Missing Drop + Typing + Explode) ──
def clean_and_type_sample(self, df: DataFrame, sample_name: str) -> DataFrame:
"""Executa limpeza de nulos 100% e tipagem inteligente por amostra.
Args:
df (DataFrame): A amostra já extraída da root_df.
sample_name (str): Nome de inferência/tabela (ex: tb_spl_stg1).
Returns:
DataFrame: Amostra purificada e tipada.
"""
import math
total_rows = df.count()
if total_rows == 0:
print(f" └── Amostra '{sample_name}' vazia, pulando...")
return df
print(f"\n{'='*72}")
print(f" PROCESSANDO INDIVIDUALMENTE: {sample_name} ({total_rows:,} linhas)")
print(f"{'='*72}")
# ── 1. Drop 100% Missing columns ──
print(f"\n ┌── 1. Eliminando colunas 100% nulas")
chunk_size = 50
cols = df.columns
missing_dict = {}
n_chunks = math.ceil(len(cols) / chunk_size)
for i in range(n_chunks):
chunk = cols[i * chunk_size : (i + 1) * chunk_size]
exprs = [
F.sum(F.when(F.col(f"`{c}`").isNull(), 1).otherwise(0)).alias(c)
for c in chunk
]
row = df.agg(*exprs).collect()[0]
for c in chunk:
missing_dict[c] = (row[c] / total_rows) if row[c] is not None else 1.0
cols_to_drop = [c for c, m in missing_dict.items() if m >= 1.0]
if cols_to_drop:
print(f" │ {len(cols_to_drop)} colunas eram 100% missing e foram removidas.")
df = df.drop(*cols_to_drop)
else:
print(f" │ Nenhuma coluna 100% nula detectada.")
remaining_cols = df.columns
# ── 2. Type Inference (respeitando features manuais) ──
analyzer = ColumnTypeAnalyzer(self.config.spark, self.config)
clf = analyzer.analyze(df)
self.column_classification = clf # Set globally if needed
# ── Print Tático Exigido ──
print(f"\n ┌── RELATÓRIO DO SCHEMA FINAL EM {sample_name}")
print(f" │ {'Coluna'.ljust(40)} | {'% Missing':^10} | Tipo")
print(f" │ " + "-" * 65)
for c in remaining_cols:
m_pct = missing_dict.get(c, 0.0)
# Find its computed category
if c in clf.numeric or c in clf.numeric_inferred: cat = "Numérico"
elif c in clf.flag_binary: cat = "Flag/Boleano"
elif c in clf.date_inferred or c in clf.temporal: cat = "Data/Temporal"
elif c in clf.numeric_vectorized or c in clf.categorical_vectorized: cat = "Vetorizada"
elif c in clf.categorical: cat = "Categórica"
else: cat = "Desconhecido/Complexo"
print(f" │ {c[:38].ljust(40)} | {m_pct:>8.1%} | {cat}")
# ── 3. Apply the Type Casts & Explode ──
df = self.apply_inferred_types(df=df, clf=clf, save_tag=sample_name)
df = self.explode_vectorized_columns(df=df, clf=clf, save_tag=sample_name)
# Final GC run per block
self.release_memory()
return df
# ── 7.1a Column Type Analysis & Vectorized Explosion ──────────
def analyze_column_types(self) -> ColumnClassification:
"""Executes automatic column type detection on the source data.
Analyses every column to classify it as numeric, categorical,
numeric_vectorized (``#``-separated numbers), or
categorical_vectorized (``#``-separated strings).
Results are persisted to Delta for governance.
Usage (independent)::
suite.load_source()
clf = suite.analyze_column_types()
print(clf.numeric_vectorized) # detected vectorised cols
print(clf.vector_lengths) # max element counts
Expected Input Schema:
``source_df`` must be loaded via ``load_source()``.
Output Schema Transformation:
Creates Delta table:
``{dest_schema}.tb_column_classification_{cart}_{YYYYMM}``.
Stores result in ``self.column_classification``.
Memory Implications:
See ``ColumnTypeAnalyzer.analyze()``.
Exception Handling Protocols:
Auto-calls ``load_source()`` if not yet done.
Propagates from ``ColumnTypeAnalyzer``.
Returns:
ColumnClassification: Full classification result.
"""
if self.source_df is None:
self.load_source()
# Try to recover classification from Delta Checkpoint
recovered = self._recover_classification_from_delta()
if recovered is not None:
self.column_classification = recovered
return self.column_classification
analyzer = ColumnTypeAnalyzer(self.config.spark, self.config)
self.column_classification = analyzer.analyze(self.source_df)
return self.column_classification
def explode_vectorized_columns(self, df: Optional[DataFrame] = None, clf: Optional['ColumnClassification'] = None, save_tag: str = "source") -> DataFrame:
"""Explodes detected vectorised columns into sub-columns.
Must be called AFTER ``analyze_column_types()``. Creates new
sub-columns ``{col}_{i}`` for each vectorised column while
**preserving** the originals.
Usage (independent)::
suite.load_source()
suite.analyze_column_types()
df = suite.explode_vectorized_columns()
print(len(df.columns)) # includes new sub-columns
Expected Input Schema:
``source_df`` loaded AND ``column_classification`` populated.
Output Schema Transformation:
Adds sub-columns to ``source_df``. Re-caches the result.
Memory Implications:
See ``VectorizedColumnHandler.explode_all()``.
Exception Handling Protocols:
Skips if ``enable_vectorized_explosion = False``.
Auto-runs ``analyze_column_types()`` if not yet done.
Returns:
DataFrame: Source DataFrame with exploded sub-columns.
"""
use_df = df if df is not None else self.source_df
if self.config.enable_delta_checkpoints:
cart_slug = self.config.carteira_ifrs.lower().replace(" ", "_")
table_name = f"{self.config.checkpoint_prefix}exploded_{save_tag}_{cart_slug}"
full_path = f"{self.config.dest_schema}.{table_name}"
try:
chkpt_df = self.config.spark.table(full_path)
print(f" ✔ Checkpoint exploded encontrado: {full_path} (Bypass!)")
chkpt_df = chkpt_df.localCheckpoint(eager=False)
chkpt_df.persist(self._get_storage_level())
if df is None: self.source_df = chkpt_df
return chkpt_df
except Exception:
pass
if not self.config.enable_vectorized_explosion:
print(" ├── Explosão de vetorizadas desabilitada (config).")
return use_df
use_clf = clf if clf is not None else self.column_classification
if use_clf is None:
use_clf = self.analyze_column_types()
n_vec = (
len(use_clf.numeric_vectorized)
+ len(use_clf.categorical_vectorized)
)
if n_vec == 0:
print(" ├── Nenhuma coluna vetorizada detectada.")
return use_df
# Unpersist old cache before transformation
use_df.unpersist()
gc.collect()
handler = VectorizedColumnHandler(
self.config.spark, self.config,
)
use_df = handler.explode_all(
use_df, use_clf,
)
# Re-persist with spill safety
if self.config.enable_delta_checkpoints:
use_df = self._checkpoint_dataframe(use_df, f"exploded_{save_tag}")
else:
sl = self._get_storage_level()
use_df.persist(sl)
if df is None: self.source_df = use_df
n = use_df.count()
print(
f" ✔ DataFrame re-persistido: {n:,} registros × "
f"{len(use_df.columns)} colunas"
)
gc.collect()
return use_df
def apply_inferred_types(self, df: Optional[DataFrame] = None, clf: Optional['ColumnClassification'] = None, save_tag: str = "source") -> DataFrame:
"""Applies automatic type casting based on column classification.
Casts ``numeric_inferred`` columns from string to ``double``
and ``date_inferred`` columns from string to ``date`` in the
source DataFrame. Uses a single ``df.select(...)`` pass to
avoid deep Catalyst plan nesting.
Must be called AFTER ``analyze_column_types()``.
Usage (independent)::
suite.load_source()
suite.analyze_column_types()
suite.apply_inferred_types()
Expected Input Schema:
``source_df`` loaded AND ``column_classification`` populated.
Output Schema Transformation:
- ``numeric_inferred`` columns: ``string`` → ``double``.
- ``date_inferred`` columns: ``string`` → ``date``.
- All other columns: unchanged.
Re-caches the result.
Memory Implications:
Single ``select`` pass. Re-caches.
Exception Handling Protocols:
Auto-calls ``analyze_column_types()`` if not yet done.
Columns that fail casting produce nulls (Spark default).
Returns:
DataFrame: Source DataFrame with inferred types applied.
"""
use_df = df if df is not None else self.source_df
if self.config.enable_delta_checkpoints:
cart_slug = self.config.carteira_ifrs.lower().replace(" ", "_")
table_name = f"{self.config.checkpoint_prefix}typed_{save_tag}_{cart_slug}"
full_path = f"{self.config.dest_schema}.{table_name}"
try:
chkpt_df = self.config.spark.table(full_path)
print(f" ✔ Checkpoint typed encontrado: {full_path} (Bypass!)")
chkpt_df = chkpt_df.localCheckpoint(eager=False)
chkpt_df.persist(self._get_storage_level())
if df is None: self.source_df = chkpt_df
return chkpt_df
except Exception:
pass
use_clf = clf if clf is not None else self.column_classification
if use_clf is None:
use_clf = self.analyze_column_types()
n_num = len(use_clf.numeric_inferred)
n_date = len(use_clf.date_inferred)
if n_num == 0 and n_date == 0:
print(" ├── Nenhuma coluna para retipagem automática.")
return use_df
print(f"\n ┌── RETIPAGEM AUTOMÁTICA")
print(f" │ Numéricas (string→double): {n_num}")
print(f" │ Datas (string→date): {n_date}")
num_set = set(use_clf.numeric_inferred)
date_set = set(use_clf.date_inferred)
# Build select expressions in a single pass
select_exprs = []
for c in use_df.columns:
cr = F.col(f"`{c}`")
if c in num_set:
select_exprs.append(
F.trim(cr).cast("double").alias(c)
)
elif c in date_set:
select_exprs.append(
F.to_date(F.trim(cr)).alias(c)
)
else:
select_exprs.append(cr)
# Unpersist old cache
use_df.unpersist()
gc.collect()
use_df = use_df.select(*select_exprs)
# Truncate lineage to prevent StackOverflow
use_df = use_df.localCheckpoint(eager=True)
if self.config.enable_delta_checkpoints:
use_df = self._checkpoint_dataframe(use_df, f"typed_{save_tag}")
else:
sl = self._get_storage_level()
use_df.persist(sl)
if df is None: self.source_df = use_df
n = use_df.count()
# Print schema changes for validation
for c in use_clf.numeric_inferred[:10]:
print(f" │ ✔ {c}: string → double")
if n_num > 10:
print(f" │ ... e mais {n_num - 10} colunas numéricas")
for c in use_clf.date_inferred[:10]:
print(f" │ ✔ {c}: string → date")
if n_date > 10:
print(f" │ ... e mais {n_date - 10} colunas de data")
print(
f" └── ✔ Retipagem completa: {n:,} registros × "
f"{len(use_df.columns)} colunas"
)
gc.collect()
return use_df
# ── 7.1b Compute Minimum Sample Size ──────────────────────────────
def compute_minimum_sample_size(self) -> Dict[str, Any]:
"""
Calculates and prints the probabilistic minimum sample size
required to represent the filtered population according to
Cochran's formula (with FPC) and Neyman stratified adjustment.
Should be called AFTER ``load_source()`` and BEFORE
``generate_all_samples()`` to inform the user whether the
configured ``total_sample_size`` is statistically sufficient.
Expected Input Schema:
``source_df`` must be loaded.
Output Schema Transformation:
Returns a dict with minimum sizes and per-stage breakdown.
Prints a formatted report to stdout.
Memory Implications:
Two aggregations (global + per-stage). Negligible.
Exception Handling Protocols:
Returns conservative estimates if variance or mean is zero.
Returns:
Dict[str, Any]: Minimum sample size report.
"""
if self.source_df is None:
self.load_source()
report = compute_minimum_sample_size(
df=self.source_df,
target_col=self.config.target_col,
stage_col=self.config.stage_col,
confidence_level=self.config.confidence_level,
margin_of_error=self.config.margin_of_error,
)
# Advisory check against configured budget
n_recommended = report["n_stratified_neyman"]
budget = self.config.total_sample_size
if budget < n_recommended:
print(
f" ⚠ ATENÇÃO: total_sample_size ({budget:,}) é MENOR que o "
f"mínimo estatístico recomendado ({n_recommended:,}). "
f"Considere aumentar o budget para garantir representatividade."
)
else:
print(
f" ✔ total_sample_size ({budget:,}) ATENDE o mínimo "
f"estatístico ({n_recommended:,})."
)
return report
# ── 7.1c Data Quality Profiling ───────────────────────────────────
def run_data_quality_profiling(self) -> DataFrame:
"""
Executes the chunked Data Quality Profiler on the source data.
Creates a ``DataQualityProfiler`` instance, runs profiling on
all 500+ columns in OOM-safe chunks, and persists results to
Delta. The profiling DataFrame is stored in ``self.profiling_df``
for use by subsequent DQ-aware methods.
Can be called independently::
suite = IFRS9SamplerSuite(config)
suite.load_source()
profiling = suite.run_data_quality_profiling()
profiling.display()
Expected Input Schema:
``source_df`` must be loaded via ``load_source()``.
Output Schema Transformation:
Creates Delta table:
``prd.sand_crc_estudos_ifrs9.tb_dq_profiling_<cart>_<YYYYMM>``
Memory Implications:
See ``DataQualityProfiler.run_profiling()``.
Exception Handling Protocols:
Propagates from DataQualityProfiler.
Returns:
DataFrame: Profiling results.
"""
if self.source_df is None:
self.load_source()
self.profiler = DataQualityProfiler(self.config.spark, self.config)
self.profiling_df = self.profiler.run_profiling(self.source_df)
return self.profiling_df
# ── 7.2 Sample A1: Neyman 30-Strata (Stage × LGD Decil) ──────────
def _sample_a1_neyman_decil(self) -> DataFrame:
"""
Sample A1: 30 strata via Stage × decil(lgd_pond).
Neyman allocation weighted by variance of ``target_col`` within
each of the 30 cross-product strata.
Expected Input Schema:
``source_df`` with ``stage``, ``lgd_pond``, ``target_col``.
Output Schema Transformation:
Adds ``stratum_id``, ``_vol_decile``.
Memory Implications:
One Window (ntile) + one groupBy + one sampleBy.
Exception Handling Protocols:
Falls back to floor=1 if budget < n_strata × floor.
Returns:
DataFrame: Neyman-optimal sample with 30 strata.
"""
print(f"\n ── Sample A1: Neyman 30-Strata (Stage × LGD Decil) ──")
return _compute_neyman_sample(
df=self.source_df,
group_cols=[self.config.stage_col],
target_col=self.config.target_col,
total_budget=self.config.total_sample_size,
min_per_stratum=self.config.min_stratum_size,
use_ntile=True,
ntile_col=self.config.decil_col,
n_tiles=10,
)
# ── 7.3 Sample A2: Neyman Stage × id_lgd_segment ─────────────────
def _sample_a2_neyman_lgdseg(self) -> DataFrame:
"""
Sample A2: Strata = Stage × id_lgd_segment (categorical cross).
No deciles. The categorical values of ``id_lgd_segment`` within
each ``stage`` form the strata organically.
Expected Input Schema:
``source_df`` with ``stage``, ``id_lgd_segment``, ``target_col``.
Output Schema Transformation:
Adds ``stratum_id``.
Memory Implications:
One groupBy on (stage, id_lgd_segment). Stratum count depends
on cardinality of ``id_lgd_segment`` (typically < 20).
Exception Handling Protocols:
Same as A1.
Returns:
DataFrame: Neyman-optimal sample by LGD segment.
"""
print(f"\n ── Sample A2: Neyman Stage × id_lgd_segment ──")
return _compute_neyman_sample(
df=self.source_df,
group_cols=[self.config.stage_col, "id_lgd_segment"],
target_col=self.config.target_col,
total_budget=self.config.total_sample_size,
min_per_stratum=self.config.min_stratum_size,
use_ntile=False,
)
# ── 7.4 Sample A3: Neyman Stage × ead_cli ────────────────────────
def _sample_a3_neyman_ead(self) -> DataFrame:
"""
Sample A3: Strata = Stage × ead_cli quantile buckets.
Since ``ead_cli`` is continuous, we first discretise it into
5 quantile buckets (quintiles) to form categorical strata,
then apply the Neyman allocation.
Expected Input Schema:
``source_df`` with ``stage``, ``ead_cli``, ``target_col``.
Output Schema Transformation:
Adds ``_ead_bucket`` (IntegerType) and ``stratum_id``.
Memory Implications:
One Window (ntile) + one groupBy + one sampleBy.
Exception Handling Protocols:
Same as A1.
Returns:
DataFrame: Neyman-optimal sample by EAD bucket.
"""
print(f"\n ── Sample A3: Neyman Stage × ead_cli ──")
# Discretise ead_cli into quintiles within each stage
w_ead = Window.partitionBy(self.config.stage_col).orderBy("ead_cli")
df_bucketed = self.source_df.withColumn(
"_ead_bucket", F.ntile(5).over(w_ead),
)
return _compute_neyman_sample(
df=df_bucketed,
group_cols=[self.config.stage_col, "_ead_bucket"],
target_col=self.config.target_col,
total_budget=self.config.total_sample_size,
min_per_stratum=self.config.min_stratum_size,
use_ntile=False,
)
# ── 7.5 Sample B: Migration-Active Oversampling ──────────────────
def _sample_b_migration_active(self) -> DataFrame:
"""
Sample B: Oversamples contracts that migrated stage between
consecutive reporting periods.
Algorithm:
1. Compute per-contract ``lag(stage)`` ordered by
``reporting_date``.
2. Flag rows where ``stage != lag_stage`` as ``migrated=1``.
3. Draw 100% of migrated contracts (census).
4. Sub-sample stable contracts (``migrated=0``) using a
fraction = ``1 / migration_oversample_ratio``.
5. Union both sets.
The resulting sample dramatically over-represents the directional
transitions that drive ``output_provision_amount`` variation,
making Feature Importance analyses reveal migration-sensitive
variables.
Expected Input Schema:
``source_df`` with ``contract_id_orig``, ``reporting_date``,
``stage``.
Output Schema Transformation:
Adds ``_lag_stage`` (IntegerType), ``_migrated`` (IntegerType).
Memory Implications:
One Window (lag) partitioned by contract sorted by date.
Shuffle bounded by contract cardinality.
Exception Handling Protocols:
First observation per contract (no lag) is treated as stable.
Returns:
DataFrame: Migration-oversampled sample.
"""
print(f"\n ── Sample B: Migração Ativa (Oversampling) ──")
w_contract = Window.partitionBy("contract_id_orig").orderBy("reporting_date")
df_lagged = self.source_df.withColumn(
"_lag_stage", F.lag(self.config.stage_col).over(w_contract),
).withColumn(
"_migrated",
F.when(
(F.col("_lag_stage").isNotNull())
& (F.col(self.config.stage_col) != F.col("_lag_stage")),
F.lit(1),
).otherwise(F.lit(0)),
)
# Census on migrated contracts
migrated = df_lagged.filter(F.col("_migrated") == 1)
stable = df_lagged.filter(F.col("_migrated") == 0)
n_migrated = migrated.count()
n_stable = stable.count()
# Sub-sample stable contracts
stable_fraction = 1.0 / self.config.migration_oversample_ratio
stable_sampled = stable.sample(
withReplacement=False,
fraction=min(stable_fraction, 1.0),
seed=42,
)
result = migrated.unionByName(stable_sampled, allowMissingColumns=True)
n_result = result.count()
print(
f" ├── Migraram: {n_migrated:,} (100% censo) "
f"| Estáveis: {n_stable:,} × {stable_fraction:.1%} "
f"= {n_result:,} total"
)
return result
# ── 7.6 Samples C/D/E: Stage-Isolated (DQ-Aware) ──────────────────
def _sample_stage_isolated(
self, stage_value: int, apply_dq_filter: bool = True,
) -> DataFrame:
"""
Generates a sample containing EXCLUSIVELY data from a single
IFRS 9 Stage, optionally pruning columns that exceed the
configured missing-value threshold.
When ``apply_dq_filter=True`` and profiling data is available
(via ``run_data_quality_profiling()``), columns with
``missing_pct > dq_missing_threshold`` **for that specific
stage** are automatically dropped. This ensures the
``model_experimentation_suite`` receives only usable features.
Args:
stage_value (int): IFRS Stage to isolate (1, 2, or 3).
apply_dq_filter (bool): If True, drops columns exceeding
the missing threshold. If False, returns all columns.
Expected Input Schema:
``source_df`` with ``stage``. Optionally ``profiling_df``.
Output Schema Transformation:
Returns a filtered subset. When DQ filter is applied,
some columns may be dropped.
Memory Implications:
Size is proportional to the stage population within the
carteira. Stage 1 is typically 70-85% of the portfolio.
Column pruning reduces downstream memory.
Exception Handling Protocols:
Returns empty DataFrame with a warning if no data exists.
Falls back to full-column mode if profiling is unavailable.
Returns:
DataFrame: Population for the requested stage, optionally
with low-quality columns removed.
"""
print(f"\n ── Sample Stage {stage_value}: Isolado (DQ-Aware) ──")
df_stage = self.source_df.filter(
F.col(self.config.stage_col) == stage_value
)
n = df_stage.count()
print(f" ├── Stage {stage_value}: {n:,} registros (censo completo)")
if n == 0:
print(f" ⚠ Nenhum registro para Stage {stage_value}!")
return df_stage
# ── DQ-Based Feature Selection ──
if apply_dq_filter and self.profiling_df is not None and self.profiler is not None:
usable_cols = self.profiler.get_usable_columns(
self.profiling_df, stage_value,
)
if usable_cols:
# Always keep structural columns
structural = {self.config.stage_col, "reporting_date",
"contract_id_orig", "carteira_ifrs"}
keep_cols = list(structural.union(set(usable_cols)))
# Only keep columns that actually exist in the DataFrame
keep_cols = [c for c in keep_cols if c in df_stage.columns]
dropped_count = len(df_stage.columns) - len(keep_cols)
df_stage = df_stage.select(*keep_cols)
print(
f" ├── DQ Filter: manteve {len(keep_cols)} colunas, "
f"descartou {dropped_count} (missing > "
f"{self.config.dq_missing_threshold*100:.0f}%)"
)
else:
print(" ├── DQ Filter: nenhuma coluna usável encontrada — mantendo todas.")
elif apply_dq_filter:
print(
" ├── DQ Filter solicitado mas profiling indisponível. "
"Execute run_data_quality_profiling() primeiro."
)
return df_stage
# ── 7.6b Directed Migration Oversampling (B1: 1→2, B2: 2→3) ──────
def _build_lagged_df(self) -> DataFrame:
"""Builds a DataFrame with ``_lag_stage`` for migration analysis.
Computes ``lag(stage)`` over ``contract_id_orig`` ordered by
``reporting_date``. This is a shared step for B, B1, and B2.
Expected Input Schema:
``source_df`` with ``contract_id_orig``, ``reporting_date``,
``stage``.
Output Schema Transformation:
Adds ``_lag_stage`` (IntegerType) column.
Memory Implications:
One Window (lag) partitioned by contract sorted by date.
Shuffle bounded by contract cardinality.
Exception Handling Protocols:
First observation per contract has _lag_stage = null.
Returns:
DataFrame: Source with ``_lag_stage`` column.
"""
w_contract = Window.partitionBy("contract_id_orig").orderBy("reporting_date")
return self.source_df.withColumn(
"_lag_stage", F.lag(self.config.stage_col).over(w_contract),
)
def _sample_directed_migration(
self, from_stage: int, to_stage: int,
) -> DataFrame:
"""Oversamples contracts that migrated from one stage to another.
Algorithm:
1. Compute ``lag(stage)`` per contract sorted by reporting_date.
2. Filter rows where ``lag_stage == from_stage`` AND
``current_stage == to_stage`` → these are the transitions.
3. Take 100% census of transition rows.
4. Sub-sample the non-transition rows at
``1 / migration_oversample_ratio``.
5. Union both sets.
Args:
from_stage (int): Origin stage (e.g., 1).
to_stage (int): Destination stage (e.g., 2).
Expected Input Schema:
``source_df`` with ``contract_id_orig``, ``reporting_date``,
``stage``.
Output Schema Transformation:
Adds ``_lag_stage`` (IntegerType), ``_transition_type``
(StringType, e.g. "STG1_TO_STG2").
Memory Implications:
One Window (lag) partitioned by contract sorted by date.
Shuffle bounded by contract cardinality. Census on
transition rows + sub-sample on remainder.
Exception Handling Protocols:
Returns empty DataFrame if no transitions exist.
Returns:
DataFrame: Migration-oversampled for the specific transition.
"""
label = f"Stage {from_stage} → {to_stage}"
print(f"\n ── Sample Migration: {label} ──")
df_lagged = self._build_lagged_df()
# Tag the specific transition
transition_tag = f"STG{from_stage}_TO_STG{to_stage}"
df_tagged = df_lagged.withColumn(
"_transition_type",
F.when(
(F.col("_lag_stage") == from_stage)
& (F.col(self.config.stage_col) == to_stage),
F.lit(transition_tag),
).otherwise(F.lit("STABLE")),
)
# Census on transitions
transitioned = df_tagged.filter(F.col("_transition_type") == transition_tag)
stable = df_tagged.filter(F.col("_transition_type") == "STABLE")
n_trans = transitioned.count()
n_stable = stable.count()
if n_trans == 0:
print(f" ⚠ Nenhuma transição {label} encontrada!")
return transitioned
# Sub-sample stable
stable_fraction = 1.0 / self.config.migration_oversample_ratio
stable_sampled = stable.sample(
withReplacement=False,
fraction=min(stable_fraction, 1.0),
seed=42,
)
result = transitioned.unionByName(
stable_sampled, allowMissingColumns=True,
)
n_result = result.count()
print(
f" ├── Transições ({label}): {n_trans:,} (100% censo) "
f"| Estáveis: {n_stable:,} × {stable_fraction:.1%} "
f"= {n_result:,} total"
)
return result
def _sample_b1_migration_stg1_to_2(self) -> DataFrame:
"""Sample B1: Oversamples Stage 1 → Stage 2 transitions.
Isolates initial credit quality degradation events. These
contracts moved from "Performing" to "Under-Performing" and
are critical for understanding early-warning signal variables
in Feature Importance.
Expected Input Schema:
``source_df`` with ``contract_id_orig``, ``reporting_date``,
``stage``.
Output Schema Transformation:
Adds ``_lag_stage``, ``_transition_type``.
Memory Implications:
See ``_sample_directed_migration()``.
Exception Handling Protocols:
See ``_sample_directed_migration()``.
Returns:
DataFrame: Oversampled Stage 1→2 transitions.
"""
return self._sample_directed_migration(from_stage=1, to_stage=2)
def _sample_b2_migration_stg2_to_3(self) -> DataFrame:
"""Sample B2: Oversamples Stage 2 → Stage 3 transitions.
Isolates the default flow events. These contracts moved from
"Under-Performing" to "Non-Performing" (default) and are
essential for validating loss-driven provision models.
Expected Input Schema:
``source_df`` with ``contract_id_orig``, ``reporting_date``,
``stage``.
Output Schema Transformation:
Adds ``_lag_stage``, ``_transition_type``.
Memory Implications:
See ``_sample_directed_migration()``.
Exception Handling Protocols:
See ``_sample_directed_migration()``.
Returns:
DataFrame: Oversampled Stage 2→3 transitions.
"""
return self._sample_directed_migration(from_stage=2, to_stage=3)
# ── 7.7 Generate All ─────────────────────────────────────────────
def generate_all_samples(self) -> Dict[str, DataFrame]:
"""
Orchestrates the generation of all 9 samples:
A1-A3 (Neyman), B (all migrations), B1 (1→2), B2 (2→3),
C/D/E (Stage-Isolated with DQ filter).
Expected Input Schema:
``source_df`` must be loaded via ``load_source()``.
Output Schema Transformation:
Populates ``self.samples`` dict with 9 named entries.
Memory Implications:
Each sample triggers one or more distributed operations
(groupBy, sampleBy, Window). No data materialises on the
driver except stratum statistics (~30-100 rows per sample).
Exception Handling Protocols:
Individual sample failures are caught and logged. The suite
continues generating remaining samples.
Returns:
Dict[str, DataFrame]: All generated samples by table name.
"""
if self.source_df is None:
self.load_source()
cart = self.config.carteira_ifrs.lower().replace(" ", "_")
sample_generators = {
f"tb_spl_ney_decil_{cart}": self._sample_a1_neyman_decil,
f"tb_spl_ney_lgdseg_{cart}": self._sample_a2_neyman_lgdseg,
f"tb_spl_ney_ead_{cart}": self._sample_a3_neyman_ead,
f"tb_spl_migr_actv_{cart}": self._sample_b_migration_active,
f"tb_spl_migr_stg1_to_2_{cart}": self._sample_b1_migration_stg1_to_2,
f"tb_spl_migr_stg2_to_3_{cart}": self._sample_b2_migration_stg2_to_3,
f"tb_spl_stg1_{cart}": lambda: self._sample_stage_isolated(1),
f"tb_spl_stg2_{cart}": lambda: self._sample_stage_isolated(2),
f"tb_spl_stg3_{cart}": lambda: self._sample_stage_isolated(3),
}
self.samples.clear()
for base_name, generator_fn in sample_generators.items():
try:
print(f"\n ├── Gerando amostra bruta: {base_name} ...")
sample_df = generator_fn()
# Materialize and count
n = sample_df.count()
# Append exact sample size to table name
table_name = f"{base_name}_n{n}"
full_table = f"{self.config.dest_schema}.{table_name}"
# Immediately save to Delta to physically persist the raw sample
if self.config.enable_delta_checkpoints:
print(f" │ Gravando Delta bruto: {full_table} ...")
sample_df.write.format("delta").mode("overwrite").option(
"overwriteSchema", "true",
).saveAsTable(full_table)
# Unpersist old, load from Delta to break lineage natively
sample_df.unpersist()
sample_df = self.config.spark.table(full_table)
sample_df = sample_df.localCheckpoint(eager=False)
sample_df.persist(self._get_storage_level())
self.samples[table_name] = sample_df
print(f" ✔ Amostra '{table_name}' gravada com {n} linhas.")
except Exception as e:
print(f" ✗ FALHA ao gerar '{base_name}': {e}")
print(f"\n ✔ Total de samples gerados com sucesso: {len(self.samples)}")
return self.samples
# ── 7.8 Validate All ─────────────────────────────────────────────
def validate_all(self) -> List[DataFrame]:
"""
Runs PSI + PRS validation on every generated sample.
Expected Input Schema:
``self.samples`` must be populated.
Output Schema Transformation:
Returns a list of validation DataFrames (one per sample).
Memory Implications:
Runs ``approxQuantile`` per column per sample. With ~7
samples × 3 columns = ~21 computations. Moderate.
Exception Handling Protocols:
Validation failures are logged; the suite continues.
Returns:
List[DataFrame]: Validation results per sample.
"""
print(f"\n{'='*72}")
print(f" VALIDAÇÃO ESTATÍSTICA (PSI + PRS)")
print(f"{'='*72}")
for table_name, sample_df in self.samples.items():
try:
report = validate_sample(
reference_df=self.source_df,
sample_df=sample_df,
sample_name=table_name,
config=self.config,
)
if report is not None:
self.validation_reports.append(report)
except Exception as e:
print(f" ✗ Validação falhou para '{table_name}': {e}")
return self.validation_reports
# ── 7.9 Persist All ──────────────────────────────────────────────
def persist_all(self) -> None:
"""
Writes every generated sample to Delta in the destination schema.
Uses ``mode("overwrite")`` with ``overwriteSchema=True`` for safe
sandbox re-runs. Each table is persisted as a managed Delta table
in Unity Catalog.
Expected Input Schema:
``self.samples`` must be populated.
Output Schema Transformation:
Creates/overwrites Delta tables in ``config.dest_schema``.
Memory Implications:
Triggers a full write action per sample. Disk I/O depends on
sample size. Parallelism managed by Spark's write planner.
Exception Handling Protocols:
Individual write failures are logged; the suite continues.
"""
print(f"\n{'='*72}")
print(f" PERSISTÊNCIA DELTA → {self.config.dest_schema}")
print(f"{'='*72}")
for table_name, sample_df in self.samples.items():
full_table = f"{self.config.dest_schema}.{table_name}"
try:
sample_df.write.format("delta").mode("overwrite").option(
"overwriteSchema", "true",
).saveAsTable(full_table)
n = sample_df.count()
print(f" ✔ {full_table} → {n:,} registros")
except Exception as e:
print(f" ✗ FALHA ao persistir '{full_table}': {e}")
# ── 7.10 Run All ─────────────────────────────────────────────────
def run(self) -> Dict[str, DataFrame]:
"""
Full execution: load → profile → min_size → generate → validate → persist.
This is the "run everything" convenience method. For production
clusters with memory constraints, prefer calling each step
independently (see ``run_step_by_step()`` docstring for guidance).
Expected Input Schema:
Fully-configured ``SamplerConfig``.
Output Schema Transformation:
Creates 9 Delta tables + 1 profiling table in
``config.dest_schema``.
Memory Implications:
Aggregate of all sub-steps. Heavy on wide tables (500+ cols).
Exception Handling Protocols:
Catches and reports failures at each phase.
Returns:
Dict[str, DataFrame]: All generated samples.
"""
self.load_source()
self.compute_minimum_sample_size()
# 1. Gera amostras brutas e já persiste em Delta
self.generate_all_samples()
self.release_memory()
# 2. Processamento individual de Type Inference e Missing Drops (reduz uso de cluster em 99%)
processed_samples = {}
for name, sample_df in self.samples.items():
cleaned_df = self.clean_and_type_sample(sample_df, name)
processed_samples[name] = cleaned_df
self.samples = processed_samples
self.release_memory()
# 3. Validações Estatísticas Base
self.validate_all()
self.release_memory()
# 4. Destinação Final
self.persist_all()
self.release_memory(full=True)
print(f"\n{'='*72}")
print(f" ✔ IFRS9 Sampler Suite — COMPLETO (ARQUITETURA OTIMIZADA)")
print(f" Samples Processados: {len(self.samples)}")
print(f" Relatórios de Validação: {len(self.validation_reports)}")
print(f"{'='*72}")
return self.samples
# ── 7.11 Step-by-Step Execution Guide ─────────────────────────────
@staticmethod
def run_step_by_step_guide() -> None:
"""Prints the recommended step-by-step execution guide.
For production clusters with memory constraints, run each step
in a separate Databricks notebook cell to allow Spark to release
intermediate caches between phases.
Expected Input Schema:
None.
Output Schema Transformation:
None. Prints to stdout.
Memory Implications:
None.
Exception Handling Protocols:
None.
"""
guide = """
╔═══════════════════════════════════════════════════════════════════════════╗
║ IFRS9 Sampler Suite — GUIA OTIMIZADO PARA CLUSTER (128GB/16c/6w) ║
║ Execute cada célula separadamente. Chame release_memory() entre ║
║ etapas pesadas para recuperar memória dos executors. ║
╚═══════════════════════════════════════════════════════════════════════════╝
# ── CELL 0: Spark Config (EXECUTAR PRIMEIRO, antes de qualquer código) ──
# Ajustes para cluster 128GB / 16 cores / 6 workers
spark.conf.set("spark.sql.shuffle.partitions", "96")
spark.conf.set("spark.sql.adaptive.enabled", "true")
spark.conf.set("spark.sql.adaptive.coalescePartitions.enabled", "true")
spark.conf.set("spark.sql.adaptive.skewJoin.enabled", "true")
spark.conf.set("spark.sql.codegen.wholeStage", "true")
spark.conf.set("spark.sql.codegen.maxFields", "200")
spark.conf.set("spark.memory.fraction", "0.8")
spark.conf.set("spark.memory.storageFraction", "0.3")
spark.conf.set("spark.sql.autoBroadcastJoinThreshold", "-1")
# Desabilita broadcast para evitar OOM no driver com tabelas largas
# ── CELL 1: Configuração e Load ──────────────────────────────────────────
from ifrs9_sampler_suite import SamplerConfig, IFRS9SamplerSuite
config = SamplerConfig(
spark=spark,
carteira_ifrs="Corporate",
reporting_date_start="2022-01-01",
dq_chunk_size=50,
dq_missing_threshold=0.40,
total_sample_size=50_000,
# ── Delta Checkpointing (Prevenção de Perdas) ──
enable_delta_checkpoints=True, # Salva estado no Delta a cada passo crítico
checkpoint_prefix="chkpt_", # Tabelas serão: chkpt_source_corporate, etc.
# ── Memória / cluster ──
n_repartition=96, # 16 cores × 6 workers
storage_level="MEMORY_AND_DISK", # spill para disco se memória cheia
enable_sample_collection=True, # False para pular coleta de exemplos
skip_date_inference=False, # True para pular detecção de datas
vectorized_chunk_size=20, # menor = menos pressão por agg()
)
# Se o cluster cair (ex: OOM), você pode rodar novamente desde o começo!
# O método load_source() verá as tabelas "chkpt_..." e as carregará em segundos
# em vez de refazer horas de processamento.
suite = IFRS9SamplerSuite(config)
suite.load_source()
# ── CELL 2: Dimensão e Sorteio Precoce ──────────────────────────────────
# Com as features fundamentais castadas, sorteamos PRIMEIRO para aliviar o motor
suite.compute_minimum_sample_size()
suite.generate_all_samples()
suite.release_memory()
# ── CELL 3: Tratamento de Nulos & Tipagem Semântica por Amostra ─────────
# Agora processamos apenas os ~50k registros em vez de 10 Milhões!
# Isso garante que se uma coluna for 100% missing apenas em uma amostra
# isolada ela será dropada na hora, preservando em amostras que a tem!
processed_samples = {}
for name, sample_df in suite.samples.items():
processed_samples[name] = suite.clean_and_type_sample(sample_df, name)
suite.samples = processed_samples
suite.release_memory()
# ── CELL 4: Validação Estatística ──────────────────────────────────────
suite.validate_all()
suite.release_memory()
# ── CELL 5: Profiling Opcional ──────────────────────────────────────────
# Se quiser o relatório completo de DQ na base inteira (Aviso: Uso de Driver Extremo)
# profiling = suite.run_data_quality_profiling()
# profiling.display()
# ── CELL 6: Persistência Delta Mestre ────────────────────────────────────
# As tabelas chkpt_* serão movidas para seu lugar oficial
suite.persist_all()
suite.release_memory(full=True) # Limpar TUDO após persistência
\"\"\"
print(guide)
# ╔═════════════════════════════════════════════════════════════════════════╗
# ║ 8. ENTRYPOINT ║
# ╚═════════════════════════════════════════════════════════════════════════╝
def main(
spark: SparkSession,
carteira: str = "Corporate",
reporting_date_start: Optional[str] = "2022-01-01",
run_profiling: bool = True,
dq_missing_threshold: float = 0.40,
dq_chunk_size: int = 50,
) -> Dict[str, DataFrame]:
"""
Standalone entrypoint for the IFRS 9 Sampler Suite.
Usage in Databricks notebook::
from ifrs9_sampler_suite import main
samples = main(spark, carteira="Corporate")
# Without profiling (faster, less cluster pressure):
samples = main(spark, carteira="Corporate", run_profiling=False)
# See step-by-step guide for modular execution:
from ifrs9_sampler_suite import IFRS9SamplerSuite
IFRS9SamplerSuite.run_step_by_step_guide()
Args:
spark (SparkSession): Active Spark session.
carteira (str): IFRS portfolio to sample.
reporting_date_start (Optional[str]): Temporal filter.
run_profiling (bool): If True, runs the full Data Quality
Profiler before sampling. Set to False for faster runs
or when profiling was already persisted.
dq_missing_threshold (float): Max missing% for DQ feature
selection (0-1). Default 0.40 = 40%.
dq_chunk_size (int): Columns per profiling chunk.
Expected Input Schema:
Active SparkSession with access to source and destination schemas.
Output Schema Transformation:
Creates 9 Delta tables + 1 profiling table in
``prd.sand_crc_estudos_ifrs9``.
Memory Implications:
See ``IFRS9SamplerSuite.run()``.
Exception Handling Protocols:
Propagates from ``IFRS9SamplerSuite.run()``.
Returns:
Dict[str, DataFrame]: All generated samples.
"""
config = SamplerConfig(
spark=spark,
carteira_ifrs=carteira,
reporting_date_start=reporting_date_start,
dq_missing_threshold=dq_missing_threshold,
dq_chunk_size=dq_chunk_size,
storage_level="MEMORY_AND_DISK",
)
suite = IFRS9SamplerSuite(config)
suite.load_source()
suite.analyze_column_types()
suite.release_memory()
suite.apply_inferred_types()
suite.release_memory()
suite.explode_vectorized_columns()
suite.release_memory()
if run_profiling:
suite.run_data_quality_profiling()
suite.release_memory()
suite.compute_minimum_sample_size()
suite.generate_all_samples()
suite.release_memory()
suite.validate_all()
suite.release_memory()
suite.persist_all()
suite.release_memory(full=True)
print(f"\n{'='*72}")
print(f" ✔ IFRS9 Sampler Suite — COMPLETO")
print(f" Samples: {len(suite.samples)}")
print(f" Relatórios de Validação: {len(suite.validation_reports)}")
print(f"{'='*72}")
return suite.samples